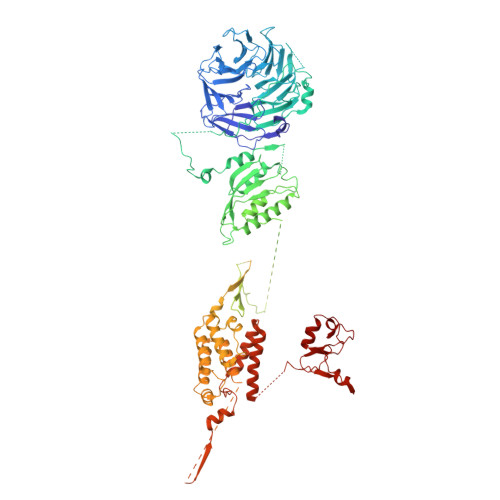
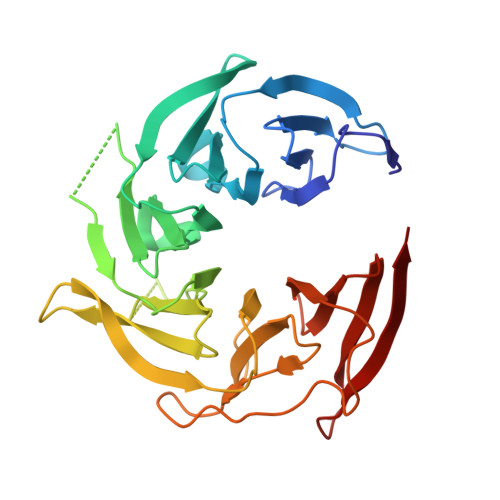
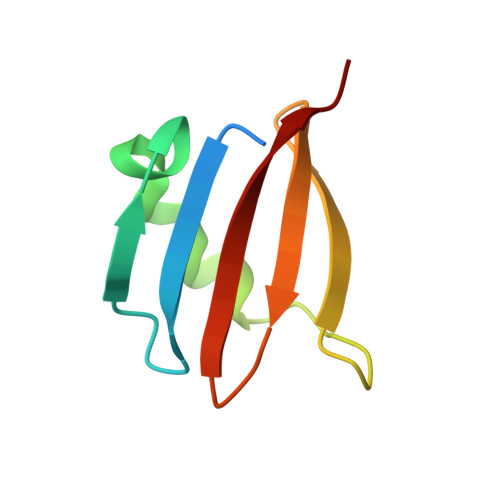
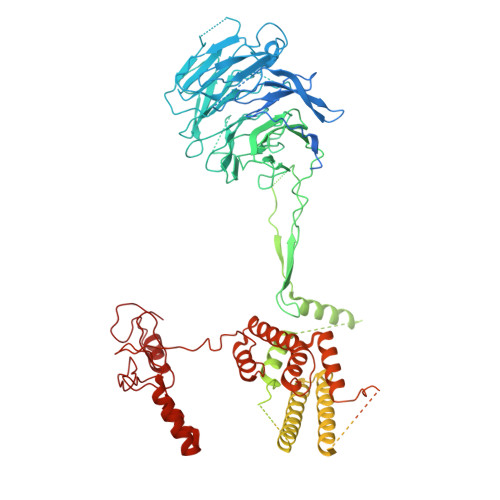
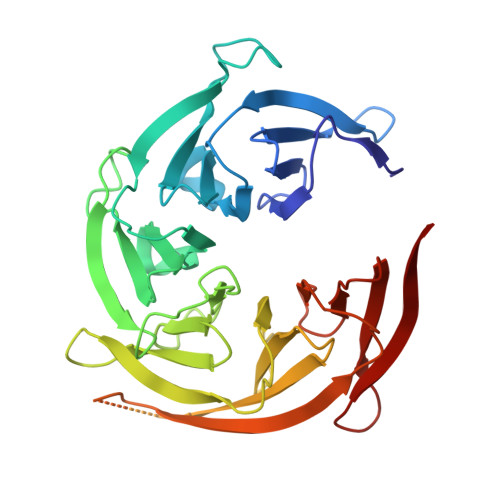
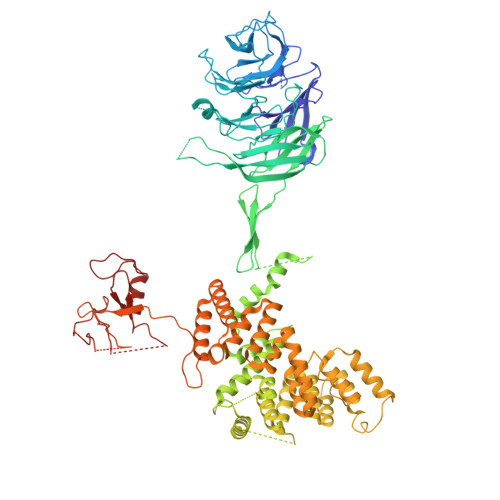
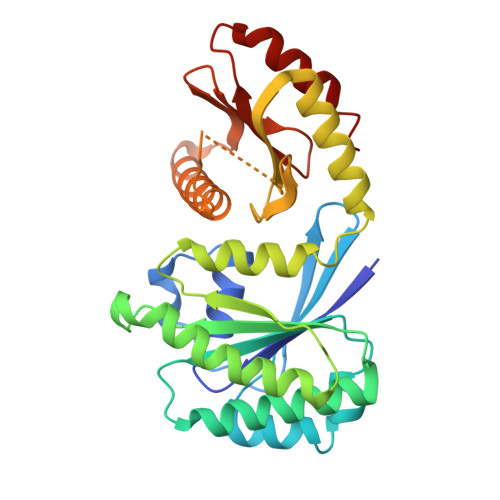
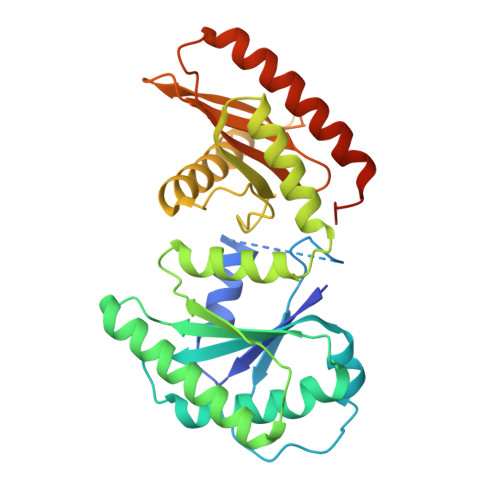
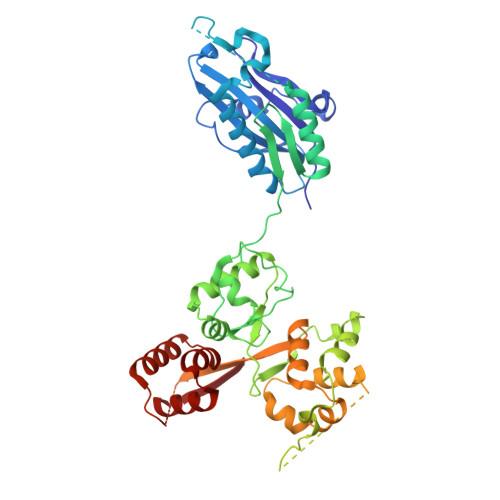
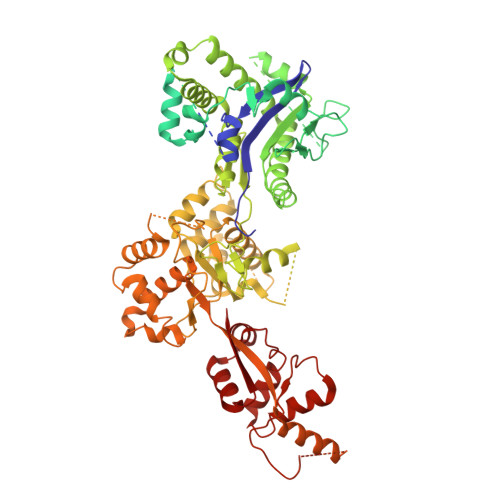
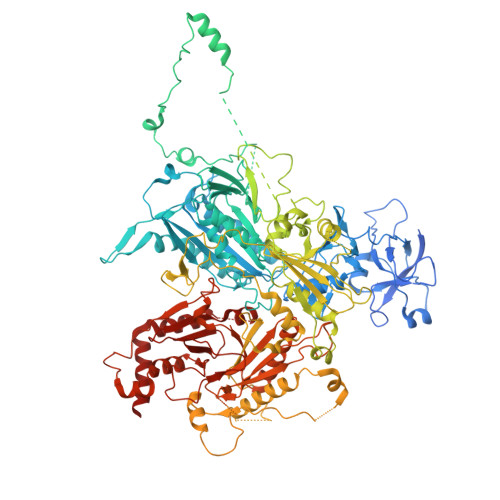
Structure and function of the yeast amino acid-sensing SEAC-EGOC supercomplex.
Tafur, L., Bonadei, L., Zheng, Y., Gabus, C., Loewith, R.(2026) Nat Struct Mol Biol
- PubMed: 41680390
- DOI: https://doi.org/10.1038/s41594-026-01746-2
- Primary Citation of Related Structures:
9H4Q, 9H5K - PubMed Abstract:
The Seh1-associated complex (SEAC; GATOR in mammals) transduces amino acid signals to the Target of Rapamycin Complex 1 (TORC1), a master regulator of cell growth. The SEAC is composed of two subcomplexes, SEACIT (GATOR1), an inhibitor of TORC1 that has GAP activity against Gtr1, and SEACAT (GATOR2), which appears to regulate SEACIT. However, the molecular details of this regulation are unclear. Here we determined the cryo-electron microscopy structure of the SEAC bound to its substrate, the EGOC (Ragulator-Rag), and studied its function in TORC1 amino acid signaling. A single SEAC can interact with two EGOC molecules via SEACIT, binding exclusively to the 'active' version of the EGOC, without involvement of SEACAT. The GAP activity of the SEACIT is essential for the regulation of TORC1 by amino acids and its loss phenocopies the lack of Gtr1-Gtr2, establishing the SEAC-EGOC complex as an amino acid-sensing hub. Compared to other SEACAT subunits, the loss of Sea2, or its N-terminal β-propeller domain, yielded strong defects in amino acid signaling to TORC1. Our results suggest that the Sea2 β-propeller recruits a GAP inhibitor to mediate fast amino acid signaling to TORC1, with additional pathways acting with slower kinetics.
- Department of Molecular and Cellular Biology, University of Geneva, Geneva, Switzerland. ltafur@cnio.es.
Organizational Affiliation: