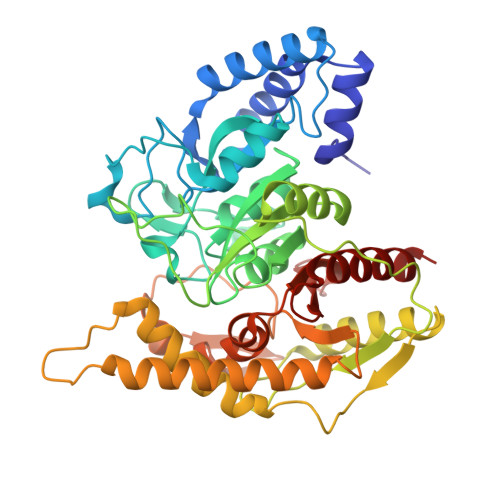
Structure-Guided Engineering of a Versatile Urethanase Improves Its Polyurethane Depolymerization Activity.
Li, Z., Han, X., Cong, L., Singh, P., Paiva, P., Branson, Y., Li, W., Chen, Y., Jaradat, D.M.M., Lennartz, F., Bayer, T., Schmidt, L., Garscha, U., You, S., Fernandes, P.A., Ramos, M.J., Bornscheuer, U.T., Weber, G., Wei, R., Liu, W.(2025) Adv Sci (Weinh) 12: e2416019-e2416019
- PubMed: 39921299
- DOI: https://doi.org/10.1002/advs.202416019
- Primary Citation of Related Structures:
8WDW, 8XTC, 9FZW - PubMed Abstract:
Polyurethane (PUR), the fifth most prevalent synthetic polymer, substantially contributes to the global plastic waste problem. Biotechnology-based recycling methods have recently emerged as innovative solutions to plastic waste disposal and sparked interest among scientific communities and industrial stakeholders in discovering and designing highly active plastic-degrading enzymes. Here, the ligand-free crystal structure of UMG-SP2, a metagenome-derived urethanase with depolymerization activities, at 2.59 Å resolution, as well as its (co-)structures bound to a suicide hydrolase inhibitor and a short-chain carbamate substrate at 2.16 and 2.40 Å resolutions, respectively, is reported. Structural analysis and molecular dynamics simulations reveal that the flexible loop L3 consisting of residues 219-226 is crucial for regulating the hydrolytic activity of UMG-SP2. The semi-rational redesign of UMG-SP2 reveals superior variants, A141G and Q399A, exhibiting over 30.7- and 7.4-fold increased activities on polyester-PUR and a methylene diamine derivative of PUR, respectively, compared to the wild-type enzyme. These findings advance the understanding of the structure-function relationship of PUR-hydrolyzing enzymes, which hold great promise for developing effective industrial PUR recycling processes and mitigating the environmental footprint of plastic waste.
- Key Laboratory of Engineering Biology for Low-carbon Manufacturing, National Engineering Research Center for Industrial Enzymes, National Technology Innovation Center of Synthetic Biology, Tianjin Institute of Industrial Biotechnology, Chinese Academy of Sciences, 32 West Seventh Avenue, Tianjin Airport Economic Area, Tianjin, 300308, China.
Organizational Affiliation: