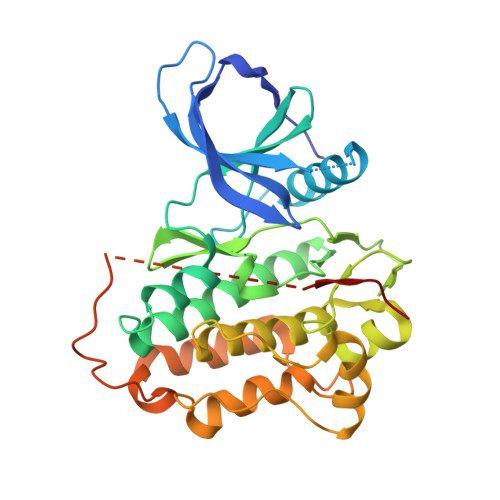
Discovery of STX-721, a Covalent, Potent, and Highly Mutant-Selective EGFR/HER2 Exon20 Insertion Inhibitor for the Treatment of Non-Small Cell Lung Cancer.
Milgram, B.C., Borrelli, D.R., Brooijmans, N., Henderson, J.A., Hilbert, B.J., Huff, M.R., Ito, T., Jackson, E.L., Jonsson, P., Ladd, B., O'Hearn, E.L., Pagliarini, R.A., Roberts, S.A., Ronseaux, S., Stuart, D.D., Wang, W., Guzman-Perez, A.(2025) J Med Chem 68: 2403-2421
- PubMed: 39824516 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.4c02377
- Primary Citation Related Structures:
9FQP, 9FQS, 9FRD - PubMed Abstract:
After L858R and ex19del epidermal growth factor receptor (EGFR) mutations, ex20ins mutations are the third most common class of driver-mutations in non-small cell lung cancer (NSCLC). Unfortunately, first-, second-, and third-generation EGFR tyrosine kinase inhibitors (TKIs) are generally ineffective for ex20ins patients due to insufficient mutant activity and selectivity over wild-type EGFR, leading to dose-limiting toxicities. While significant advances in recent years have been made toward identifying potent EGFR ex20ins mutant inhibitors, mutant vs wild-type EGFR selectivity remains a significant challenge. STX-721 ( 53 ) is a potent, irreversible inhibitor of the majority of EGFR/HER2 ex20ins mutants and demonstrates excellent mutant vs wild-type selectivity both in vitro and in vivo. STX-721 is currently in phase 1/2 clinical trials for EGFR/HER2 ex20ins-driven NSCLC.
- Scorpion Therapeutics, 1 Winthrop Square, Boston, Massachusetts 02110, United States.
Organizational Affiliation: