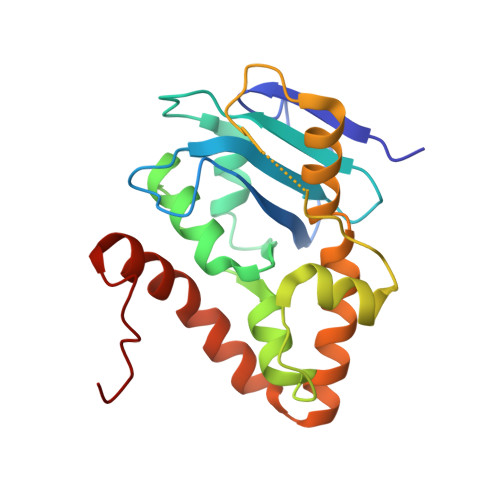
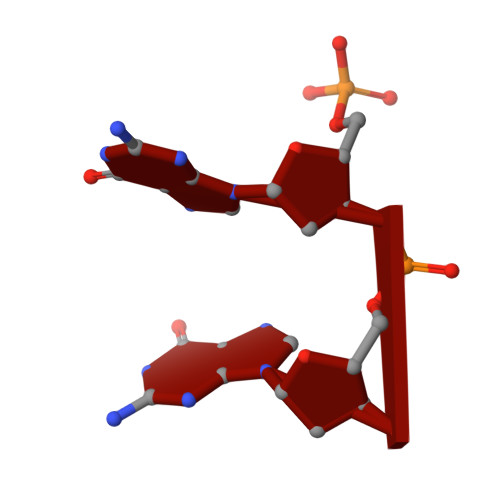
Structural and bioinformatics analyses identify deoxydinucleotide-specific nucleases and their association with genomic islands in gram-positive bacteria.
Mortensen, S., Kuncova, S., Lormand, J.D., Myers, T.M., Kim, S.K., Lee, V.T., Winkler, W.C., Sondermann, H.(2025) Nucleic Acids Res 53
- PubMed: 39778863 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkae1235
- Primary Citation Related Structures:
9F7D, 9F7G, 9F7J, 9F7L, 9F7M - PubMed Abstract:
Dinucleases of the DEDD superfamily, such as oligoribonuclease, Rexo2 and nanoRNase C, catalyze the essential final step of RNA degradation, the conversion of di- to mononucleotides. The active sites of these enzymes are optimized for substrates that are two nucleotides long, and do not discriminate between RNA and DNA. Here, we identified a novel DEDD subfamily, members of which function as dedicated deoxydinucleases (diDNases) that specifically hydrolyze single-stranded DNA dinucleotides in a sequence-independent manner. Crystal structures of enzyme-substrate complexes reveal that specificity for DNA stems from a combination of conserved structural elements that exclude diribonucleotides as substrates. Consistently, diDNases fail to complement the loss of enzymes that act on diribonucleotides, indicating that these two groups of enzymes support distinct cellular functions. The genes encoding diDNases are found predominantly in genomic islands of Actinomycetes and Clostridia, which, together with their association with phage-defense systems, suggest potential roles in bacterial immunity.
- CSSB Centre for Structural Systems Biology, Deutsches Elektronen-Synchrotron DESY, Notkestr. 85, 22607 Hamburg, Germany.
Organizational Affiliation: