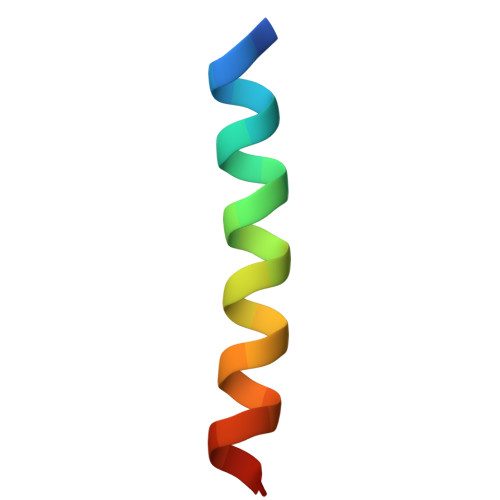
De Novo Design of Parallel and Antiparallel A 3 B 3 Heterohexameric alpha-Helical Barrels.
Chubb, J.J., Albanese, K.I., Rodger, A., Woolfson, D.N.(2025) Biochemistry 64: 1973-1982
- PubMed: 40227224 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.4c00584
- Primary Citation Related Structures:
9EVG - PubMed Abstract:
The de novo design of α-helical coiled-coil peptides is advanced. Using established sequence-to-structure relationships, it is possible to generate various coiled-coil assemblies with predictable numbers and orientations of helices. Here, we target new assemblies, namely, A 3 B 3 heterohexamer α-helical barrels. These designs are based on pairs of sequences with three heptad repeats ( abcdefg ), programmed with a = Leu, d = Ile, e = Ala, and g = Ser, and b = c = Glu to make the acidic (A) chains and b = c = Lys in the basic (B) chains. These design rules ensure that the desired oligomeric state and stoichiometry are readily achieved. However, controlling the orientation of neighboring helices (parallel or antiparallel) is less straightforward. Surprisingly, we find that assembly and helix orientation are sensitive to the length of the overhang between helices. To study this, cyclically permutated peptide sequences with three heptad repeats (the register) in the peptide sequences were analyzed. Peptides starting at g ( g -register) form a parallel 6-helix barrel in solution and in an X-ray crystal structure, whereas the b - and c -register peptides form an antiparallel complex. In lieu of experimental X-ray structures for b - and c -register peptides, AlphaFold-Multimer is used to predict atomistic models. However, considerably more sampling than the default value is required to match the models and the experimental data, as many confidently predicted and plausible models are generated with incorrect helix orientations. This work reveals the previously unknown influence of the heptad register on helical overhang and the orientation of α-helical coiled-coil peptides and provides insights for the modeling of oligopeptide coiled-coil complexes with AlphaFold.
- School of Chemistry, University of Bristol, Cantock's Close, Bristol BS8 1TS, U.K.
Organizational Affiliation: