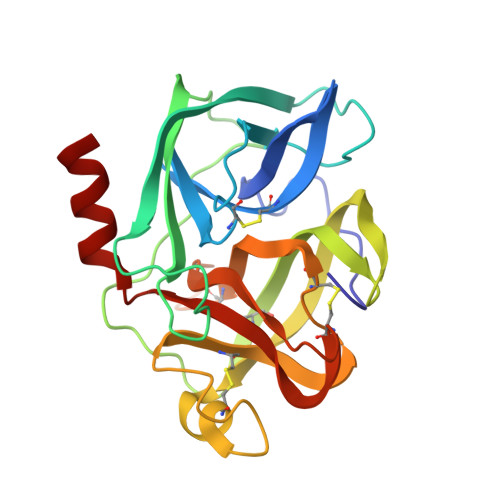
Structural study of porcine pancreatic elastase complexed with 7-amino-3-(2-bromoethoxy)-4-chloroisocoumarin as a nonreactivatable doubly covalent enzyme-inhibitor complex.
Vijayalakshmi, J., Meyer Jr., E.F., Kam, C.M., Powers, J.C.(1991) Biochemistry 30: 2175-2183
- PubMed: 1998677 Search on PubMed
- DOI: https://doi.org/10.1021/bi00222a022
- Primary Citation Related Structures:
9EST - PubMed Abstract:
The complex of porcine pancreatic elastase (PPE) with 7-amino-3-(2-bromoethoxy)-4-chloroisocoumarin, a potent mechanism-based inhibitor, was crystallized and the crystal structure determined at 1.9-A resolution with a final R factor of 17.1%. The unbiased difference Fourier electron density map showed continuous density from O gamma of Ser 195 to the benzoyl carbonyl carbon atom and from N epsilon 2 of His 57 to the carbon atom at the 4-position of the isocoumarin ring in the inhibitor. This suggested unambiguously that the inhibitor was doubly covalently bound to the enzyme. It represents the first structural evidence for irreversible binding of an isocoumarin inhibitor to PPE through both Ser 195 and His 57 in the active site. The PPE-inhibitor complex is only partially activated in solution by hydroxylamine and confirms the existence of the doubly covalently bound complex along with the acyl enzyme. The benzoyl carbonyl oxygen atom of the inhibitor is not situated in the oxyanion hole formed by the amide (greater than NH) groups of Gly 193 and Ser 195. The complex is stabilized by the hydrogen-bonding interactions in the active site (from the N epsilon 2 of Gln 192 to the bromine atom in the inhibitor and the amino group at the 7-position of the isocoumarin ring to the carbonyl oxygen of Thr 41) and by van der Waals interactions. The inhibition rates of several 7-substituted 4-chloro-3-(bromoalkoxy)isocoumarins toward PPE were measured.(ABSTRACT TRUNCATED AT 250 WORDS)
- Department of Bichemistry and Biophysics, Texas A&M University, College Station 77843-2128.
Organizational Affiliation: