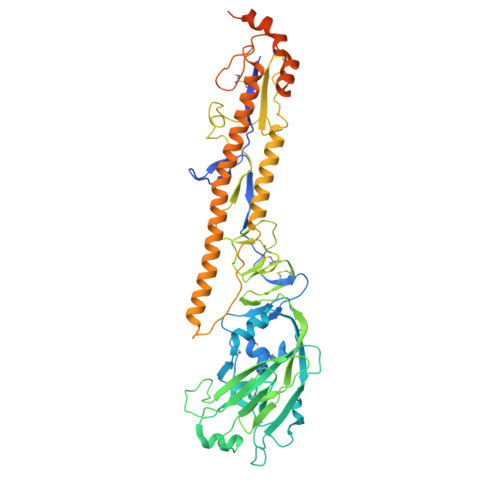
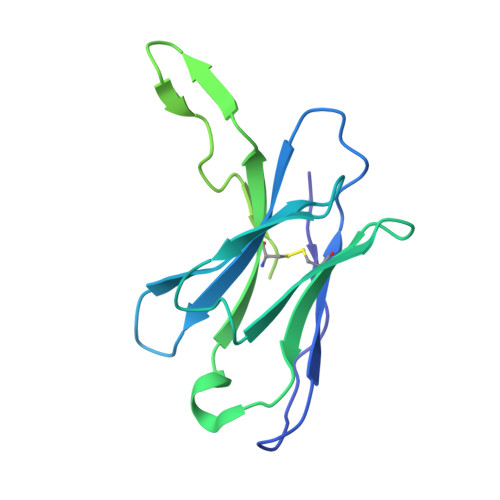
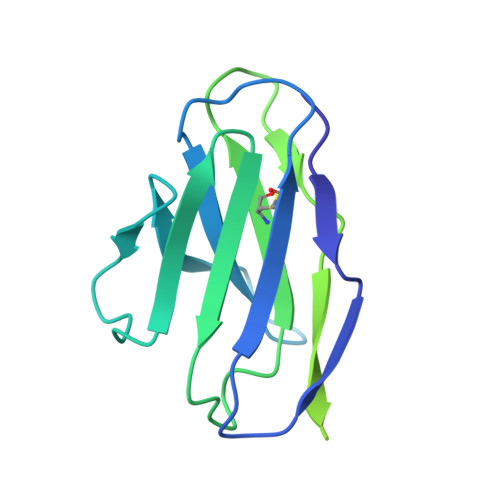
Structure of a zoonotic H5N1 hemagglutinin reveals a receptor-binding site occupied by an auto-glycan.
Morano, N.C., Guo, Y., Becker, J.E., Li, Z., Yu, J., Ho, D.D., Shapiro, L., Kwong, P.D.(2025) Structure 33: 228-233.e3
- PubMed: 39884273 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2025.01.001
- Primary Citation Related Structures:
9EKF - PubMed Abstract:
Highly pathogenic avian influenza has spilled into many mammals, most notably cows and poultry, with several dozen human breakthrough infections. Zoonotic crossovers, with hemagglutinins mutated to enhance viral ability to use human α2-6-linked sialic acid receptors versus avian α2-3-linked ones, highlight the pandemic risk. To gain insight into these crossovers, we determined the cryoelectron microscopy (cryo-EM) structure of the hemagglutinin from the zoonotic H5N1 A/Texas/37/2024 strain (clade 2.3.4.4b) in complex with a previously reported neutralizing antibody. Surprisingly, we found that the receptor-binding site of this H5N1 hemagglutinin was already occupied by an α2-3-linked sialic acid and that this glycan emanated from asparagine N169 of a neighboring protomer on hemagglutinin itself. This structure thus highlights recognition by influenza hemagglutinin of an "auto"-α2-3-linked sialic acid from N169, an N-linked glycan conserved in 95% of H5 strains, and adds "auto-glycan recognition," which may play a role in viral dispersal, to the complexities surrounding H5N1 zoonosis.
- Aaron Diamond AIDS Research Center, Columbia University Vagelos College of Physicians and Surgeons, New York, NY 10032, USA; Zuckerman Mind Brain Behavior Institute, Columbia University, New York, NY 10027, USA; Department of Biochemistry and Molecular Biophysics, Columbia University, New York, NY 10027, USA.
Organizational Affiliation: