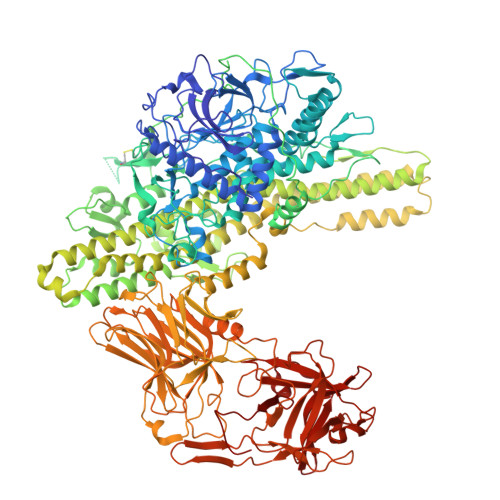
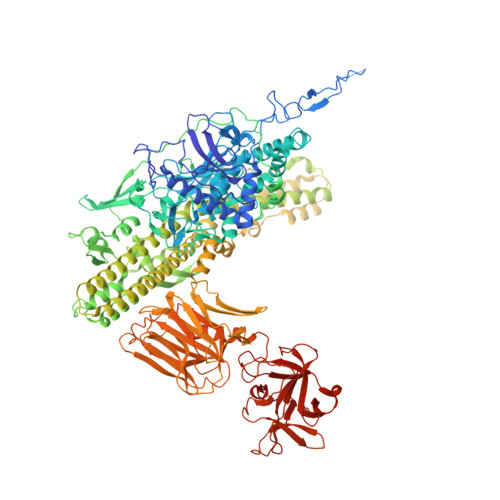
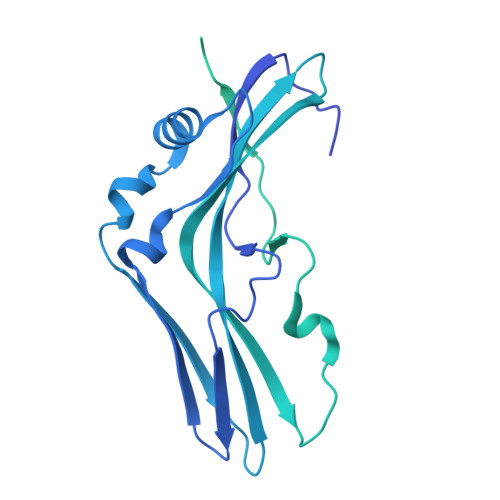
A nut-and-bolt assembly of the bimodular large progenitor botulinum neurotoxin complex.
Lam, K.H., Gao, L., Przykopanski, A., Chen, B., Huang, T., Kruger, M., Bartels, A.M., Dorner, M.B., Perry, K., Dorner, B.G., Rummel, A., Jin, R.(2025) Sci Adv 11: eadx5831
- PubMed: 40864694 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.adx5831
- Primary Citation Related Structures:
9EA9 - PubMed Abstract:
Botulinum neurotoxin serotype A (BoNT/A) is naturally produced by bacteria along with four nontoxic neurotoxin-associated proteins (NTNH, HA70, HA33, and HA17), forming a bimodular large progenitor toxin complex (L-PTC). The BoNT/A-NTNH complex protects the toxin from adverse environment, while the complex consisting of HA proteins facilitates toxin absorption during oral intoxication. How these two independent modules assemble into the L-PTC remains unclear. Here, we report the crystal structure of the BoNT/A-NTNH-HA70 complex at ~2.9-Å resolution. The structure reveals that the BoNT/A-NTNH complex is anchored into a concentric double β-barrel channel of trimeric HA70 through a short β-hairpin of NTNH (termed nLoop), resembling a nut-and-bolt attachment. We find that the nLoop of NTNH is strictly conserved across HA-containing BoNT complexes and that NTNH-HA70 binding is interchangeable among them. Furthermore, we demonstrate that the nLoop functions as a minimal motif enabling attachment of a protein-of-interest to the HA complex, with potential applications in oral biologics delivery.
- Department of Physiology and Biophysics, University of California, Irvine, CA 92697, USA.
Organizational Affiliation: