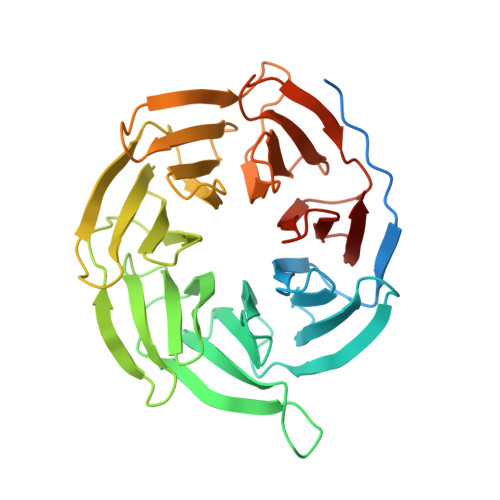
Structural basis for the activation of proteinase-activated receptors PAR1 and PAR2.
Lyu, Z., Lyu, X., Malyutin, A.G., Xia, G., Carney, D., Alves, V.M., Falk, M., Arora, N., Zou, H., McGrath, A.P., Kang, Y.(2025) Nat Commun 16: 3931-3931
- PubMed: 40287415 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-59138-x
- Primary Citation Related Structures:
9D0A, 9D4Z, 9E7R - PubMed Abstract:
Members of the proteinase-activated receptor (PAR) subfamily of G protein-coupled receptors (GPCRs) play critical roles in processes like hemostasis, thrombosis, development, wound healing, inflammation, and cancer progression. Comprising PAR1-PAR4, these receptors are specifically activated by protease cleavage at their extracellular amino terminus, revealing a 'tethered ligand' that self-activates the receptor. This triggers complex intracellular signaling via G proteins and beta-arrestins, linking external protease signals to cellular functions. To date, direct structural visualization of these ligand-receptor complexes has been limited. Here, we present structural snapshots of activated PAR1 and PAR2 bound to their endogenous tethered ligands, revealing a shallow and constricted orthosteric binding pocket. Comparisons with antagonist-bound structures show minimal conformational changes in the TM6 helix and larger movements of TM7 upon activation. These findings reveal a common activation mechanism for PAR1 and PAR2, highlighting critical residues involved in ligand recognition. Additionally, the structure of PAR2 bound to a pathway selective antagonist, GB88, demonstrates how potent orthosteric engagement can be achieved by a small molecule mimicking the endogenous tethered ligand's interactions.
- Takeda Development Center Americas, Inc, 9625 Towne Centre Drive, San Diego, CA, USA.
Organizational Affiliation: