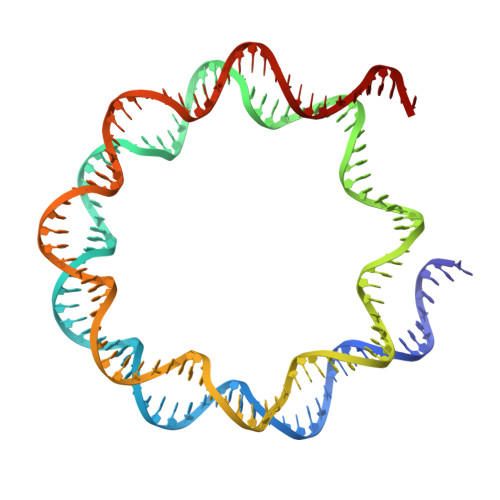
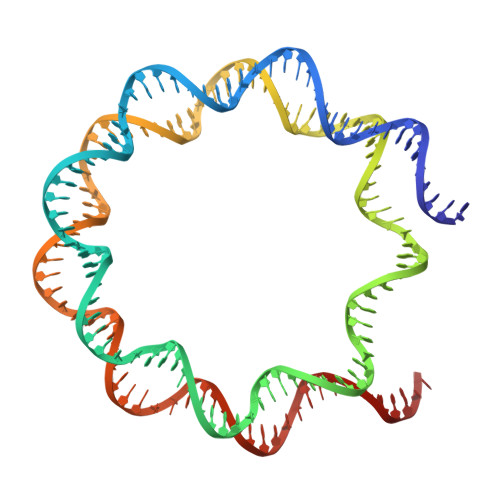
Mechanisms of chromatin remodeling by an Snf2-type ATPase.
Malik, D., Deshmukh, A., Bilokapic, S., Halic, M.(2025) bioRxiv
- PubMed: 39803580
- DOI: https://doi.org/10.1101/2024.12.31.630910
- Primary Citation Related Structures:
9E1Y - PubMed Abstract:
Chromatin remodeling enzymes play a crucial role in the organization of chromatin, enabling both stability and plasticity of genome regulation. These enzymes use a Snf2-type ATPase motor to move nucleosomes, but how they translocate DNA around the histone octamer is unclear. Here we use cryo-EM to visualize the continuous motion of nucleosomal DNA induced by human chromatin remodeler SNF2H, an ISWI family member. Our work reveals conformational changes in SNF2H, DNA and histones during nucleosome sliding and provides the structural basis for DNA translocation. ATP hydrolysis induces conformational changes in SNF2H that pull the DNA tracking strand, distorting DNA and histones at SHL2. This is followed by SNF2H rotation on the nucleosome, which first pulls the DNA guide strand and creates one-base pair bulge at SHL2, and then releases the pulled DNA. Given the high conservation of the catalytic motors among ATP-dependent chromatin remodelers, the mechanisms we describe likely apply to other families.
- Department of Structural Biology, St. Jude Children's Research Hospital, 262 Danny Thomas Place, Memphis, TN, 38105, USA.
Organizational Affiliation: