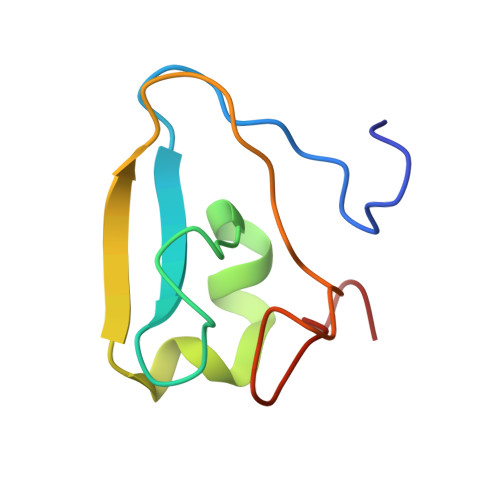
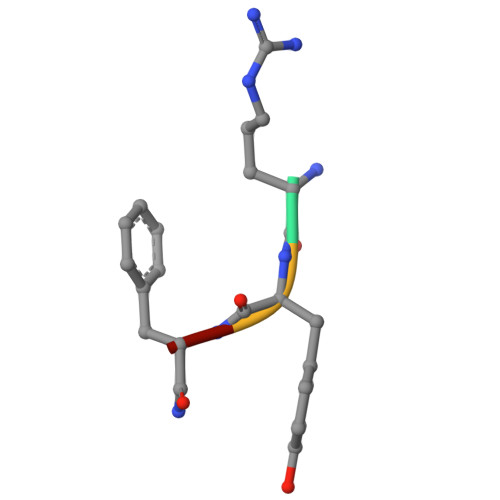
An alternative pocket for binding the N-degrons by the UBR1 and UBR2 ubiquitin E3 ligases.
Huang, S.T., Chen, D.H., Ren, T., Thomas, N., Wu, J., Sankaran, B., Jones, R., Taylor, S., Chen, Y.(2025) Protein Sci 34: e70248-e70248
- PubMed: 40880185 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.70248
- Primary Citation Related Structures:
9DNO, 9DNP, 9DNQ, 9DNR, 9MUX - PubMed Abstract:
The UBR family of ubiquitin ligases binds to N-termini of their targets (known as N-degron) to induce their ubiquitination and degradation via a conserved domain known as UBR-box. UBR1 and UBR2 share the highest sequence homology among the family, and substantial structural studies were previously performed for substrate binding by the UBR-boxes of UBR1 and UBR2. Here, we describe a new pocket in the UBR-boxes of UBR1 and UBR2 for binding the second residues of N-degrons through determining five co-crystal structures of the UBR-boxes with various N-degron peptides. Together with binding affinities measured by fluorescence polarization, we show that the two highly homologous UBR-boxes can interact with the second residue of an N-degron differently. In addition, the UBR-boxes undergo different conformational changes when binding N-degrons. Furthermore, we demonstrate that the sidechain of the third amino acid of an N-degron has no contribution to binding the UBR-boxes. These findings represent a new conceptual advancement for the UBR E3 ligases and the new insights described here can be leveraged for developing their selective ligands for research and potential therapies.
- Department of Chemistry and Biochemistry, University of California, San Diego, La Jolla, California, USA.
Organizational Affiliation: