Cryo-EM structure of the USP1-UAF1-Ubiquitin complex inhibited by KSQ-4279
Griffith, J.P., Wylie, A.A., Ali, J.A.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| WD repeat-containing protein 48 | 677 | Homo sapiens | Mutation(s): 0 Gene Names: WDR48, KIAA1449, UAF1 | 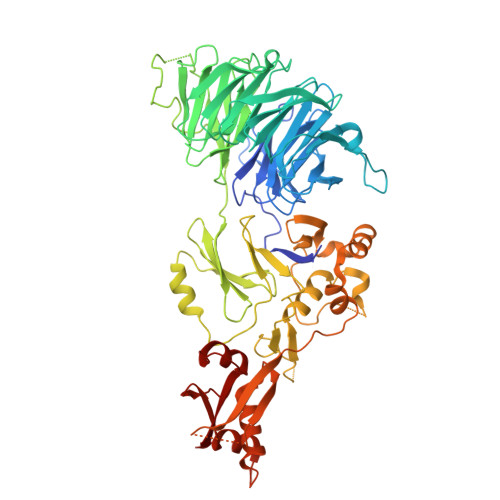 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q8TAF3 GTEx: ENSG00000114742 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q8TAF3 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Ubiquitin carboxyl-terminal hydrolase 1 | 785 | Homo sapiens | Mutation(s): 0 Gene Names: USP1 EC: 3.4.19.12 | 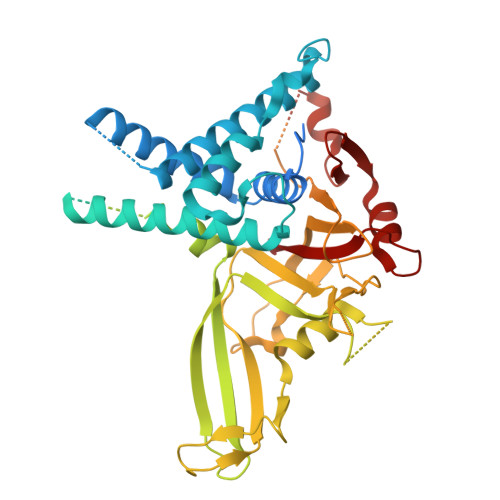 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: O94782 GTEx: ENSG00000162607 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O94782 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Ubiquitin | 75 | Homo sapiens | Mutation(s): 0 Gene Names: UBC | 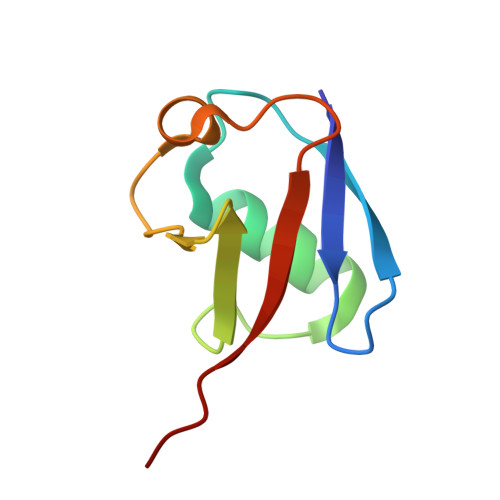 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P0CG48 GTEx: ENSG00000150991 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0CG48 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| A1IB8 Download:Ideal Coordinates CCD File | E [auth B] | 6-(4-cyclopropyl-6-methoxy-pyrimidin-5-yl)-1-[[4-[1-propan-2-yl-4-(trifluoromethyl)imidazol-2-yl]phenyl]methyl]pyrazolo[3,4-d]pyrimidine C27 H25 F3 N8 O KCBWAFJCKVKYHO-UHFFFAOYSA-N |  | ||
| A1A4Y Download:Ideal Coordinates CCD File | F [auth B] | 3-(methanesulfonyl)propan-1-amine C4 H11 N O2 S QUYFSHOLLKVPGX-UHFFFAOYSA-N |  | ||
| ZN Download:Ideal Coordinates CCD File | D [auth B] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| Funding Organization | Location | Grant Number |
|---|---|---|
| Not funded | -- |