Structure of canonical Myo7a-C isoform (ADP-bound) expressed in sensory hair cells (head domain + first two IQ domains), bound to F-actin
Egelman, E.H., Shin, J.B.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Actin, alpha skeletal muscle | 377 | Oryctolagus cuniculus | Mutation(s): 0 EC: 3.6.4 | 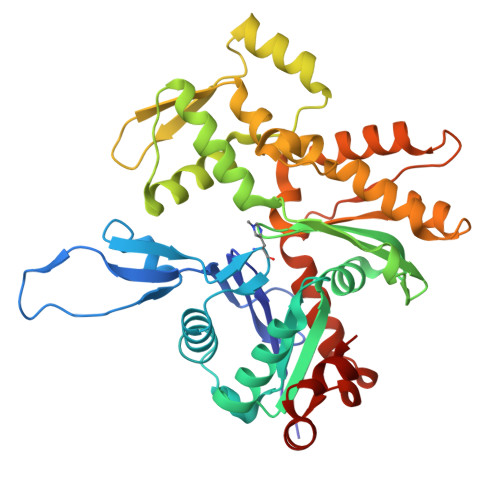 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P68135 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Unconventional myosin-VIIa | 670 | Mus musculus | Mutation(s): 0 Gene Names: Myo7a, Myo7 | 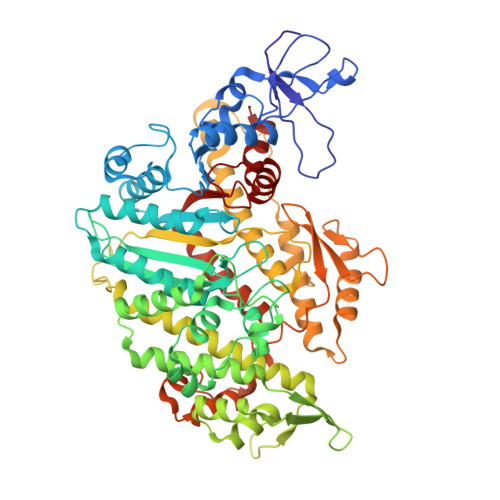 | |
UniProt & NIH Common Fund Data Resources | |||||
IMPC: MGI:104510 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P97479 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| ADP (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | E [auth A], G [auth B], I [auth C], L [auth D] | ADENOSINE-5'-DIPHOSPHATE C10 H15 N5 O10 P2 XTWYTFMLZFPYCI-KQYNXXCUSA-N |  | ||
| MG (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | F [auth A], H [auth B], J [auth C], K [auth D] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| HIC Query on HIC | A, B, C | L-PEPTIDE LINKING | C7 H11 N3 O2 |  | HIS |
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | 1.20.1_4487: |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) | United States | GM122510 |