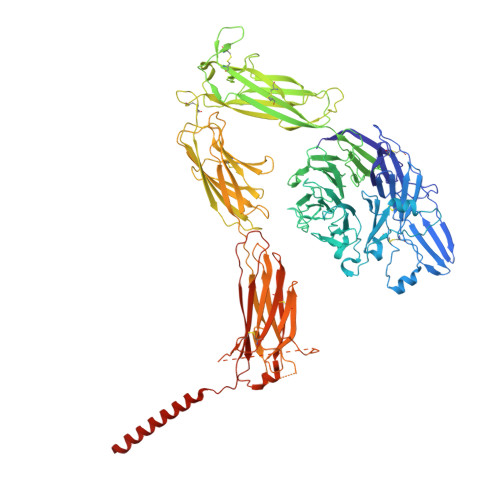
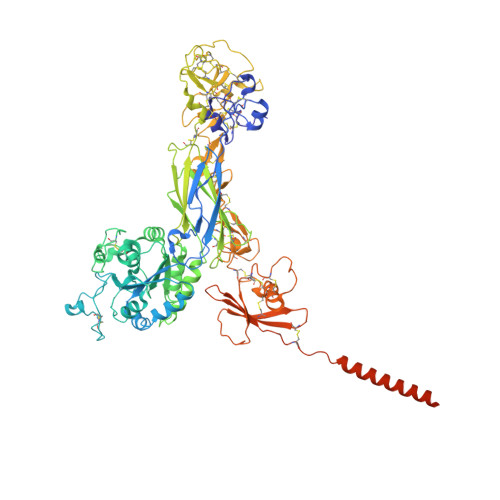
Platelet integrin alpha IIb beta 3 plays a key role in a venous thrombogenesis mouse model.
Adair, B.D., Field, C.O., Alonso, J.L., Xiong, J.P., Deng, S.X., Ahn, H.S., Mashin, E., Clish, C.B., van Agthoven, J., Yeager, M., Guo, Y., Tess, D.A., Landry, D.W., Poncz, M., Arnaout, M.A.(2024) Nat Commun 15: 8612-8612
- PubMed: 39366965 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-52869-3
- Primary Citation Related Structures:
9DEQ, 9DER - PubMed Abstract:
Venous thrombosis (VT) is a common vascular disease associated with reduced survival and a high recurrence rate. VT is initiated by the accumulation of platelets and neutrophils at sites of endothelial cell activation. A role for platelet αIIbβ3 in VT is not established, a task complicated by the increased bleeding risk caused by partial agonists such as tirofiban. Here, we show that m-tirofiban, a modified version of tirofiban, does not agonize αIIbβ3 based on lack of neoepitope expression and the cryo-EM structure of m-tirofiban/full-length αIIbβ3 complex. m-tirofiban abolishes agonist-induced platelet aggregation while preserving clot retraction ex vivo and, unlike tirofiban, it suppresses venous thrombogenesis in a mouse model without increasing bleeding. These findings establish a key role for αIIbβ3 in VT initiation and suggest that m-tirofiban and compounds with a similar structurally-defined mechanism of action merit consideration as potential thromboprophylaxis agents in patients at high risk for VT and hemorrhage.
- Division of Nephrology, Massachusetts General Hospital and Harvard Medical School, Boston, MA, USA.
Organizational Affiliation: