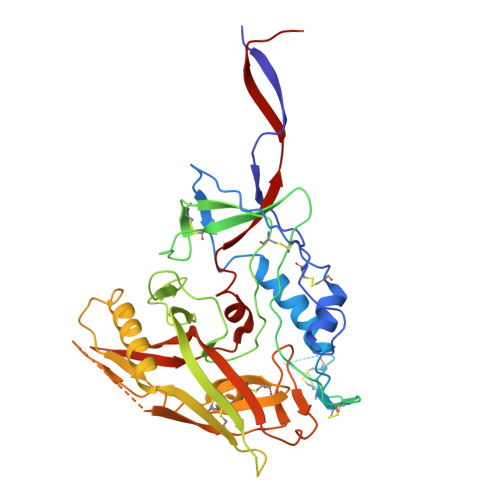
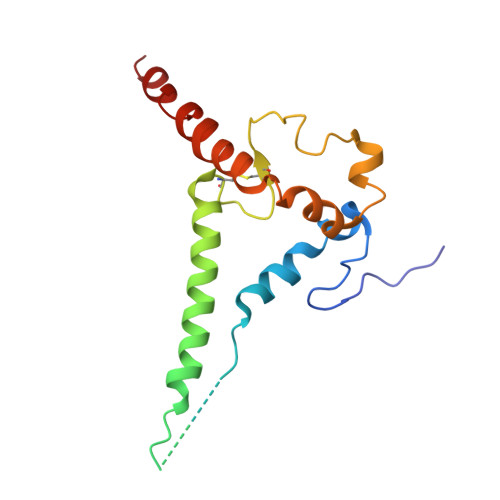
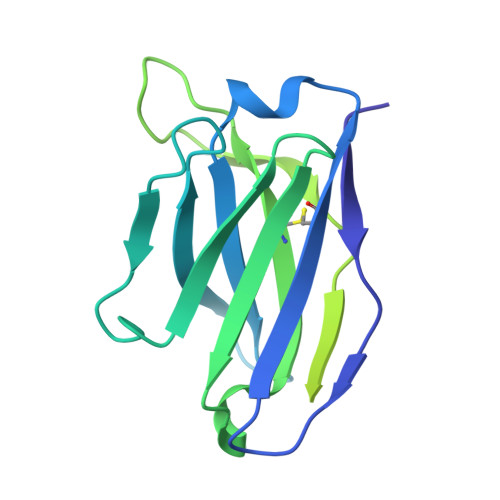
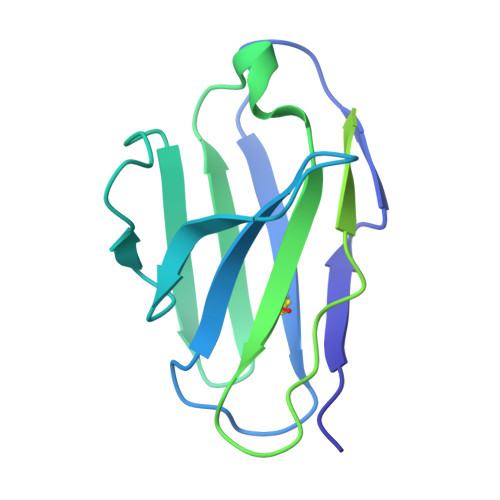
Conformational trajectory of the HIV-1 fusion peptide during CD4-induced envelope opening.
Thakur, B., Katte, R.H., Xu, W., Janowska, K., Sammour, S., Henderson, R., Lu, M., Kwong, P.D., Acharya, P.(2025) Nat Commun 16: 4595-4595
- PubMed: 40382314 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-59721-2
- Primary Citation Related Structures:
9D8Y, 9D90, 9D98 - PubMed Abstract:
The hydrophobic fusion peptide (FP), a critical component of the HIV-1 entry machinery, is located at the N terminus of the envelope (Env) gp41 subunit. The receptor-binding gp120 subunit of Env forms a heterodimer with gp41. The gp120/gp41 heterodimer assembles into a homotrimer, in which FP is accessible for antibody binding. Env conformational changes or "opening" that follow receptor binding result in FP relocating to a newly formed interprotomer pocket at the gp41-gp120 interface where it is sterically inaccessible to antibodies. The mechanistic steps connecting the entry-related transition of antibody accessible-to-inaccessible FP configurations remain unresolved. Here, using SOSIP-stabilized Env ectodomains, we visualize that the FP remains accessible for antibody binding despite substantial receptor-induced Env opening. We delineate stepwise Env opening from its closed state to a functional CD4-bound symmetrically open Env in which we show that FP was accessible for antibody binding. We define downstream re-organizations that lead to the formation of a gp120/gp41 cavity into which the FP buries to become inaccessible for antibody binding. These findings improve our understanding of HIV-1 entry and delineate the entry-related conformational trajectory of a key site of HIV vulnerability to neutralizing antibody.
- Duke Human Vaccine Institute, Duke University, Durham, NC, USA.
Organizational Affiliation: