The ancient MHC class I molecule HLA-B*38:01 calibrates the immune system to protect against multiple sclerosis
Ladell, K., Zhu, S., Kaufmann, M., Vivian, J., Petersen, J., Rossjohn, J., Price, D., Fugger, L.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| MHC class I antigen | 276 | Homo sapiens | Mutation(s): 0 Gene Names: HLA-B, HLA-DRB1 | 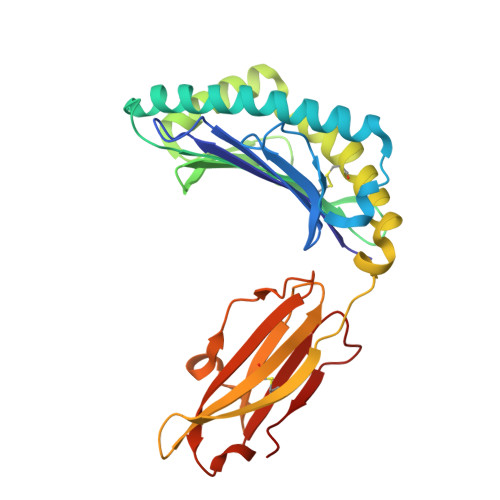 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | E5FQ58 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Beta-2-microglobulin | 100 | Homo sapiens | Mutation(s): 0 Gene Names: B2M, CDABP0092, HDCMA22P | 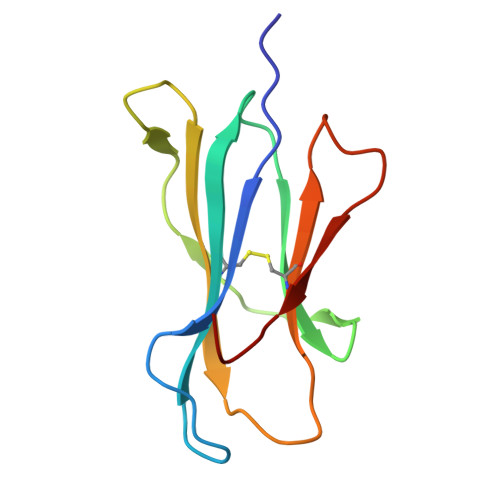 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P61769 GTEx: ENSG00000166710 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P61769 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cyclin-A2 | 9 | Homo sapiens | Mutation(s): 0 Gene Names: CCNA2, CCN1, CCNA | 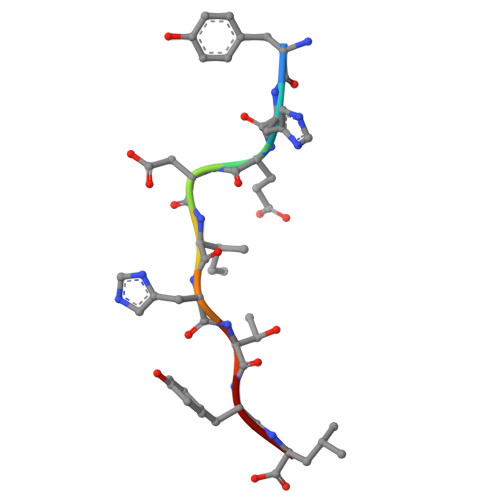 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P20248 GTEx: ENSG00000145386 | |||||
Entity Groups | |||||
| UniProt Group | P20248 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Killer cell immunoglobulin-like receptor 3DL1 | D [auth G] | 301 | Homo sapiens | Mutation(s): 0 Gene Names: KIR3DL1, CD158E, NKAT3, NKB1 | 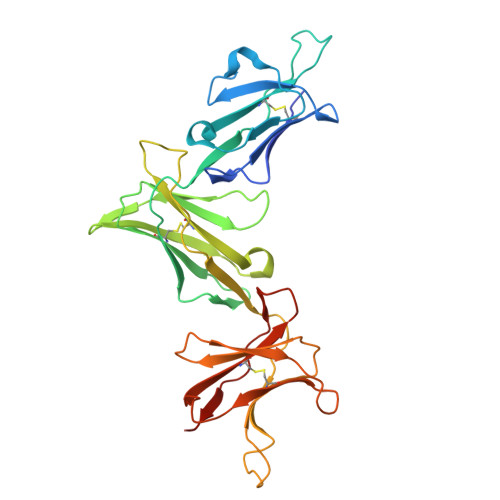 |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P43629 GTEx: ENSG00000167633 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P43629 | ||||
Glycosylation | |||||
| Glycosylation Sites: 3 | Go to GlyGen: P43629-1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAG Download:Ideal Coordinates CCD File | E [auth G], F [auth G], G | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 46.963 | α = 94.96 |
| b = 60.047 | β = 95.5 |
| c = 63.337 | γ = 106.23 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Health and Medical Research Council (NHMRC, Australia) | Australia | -- |