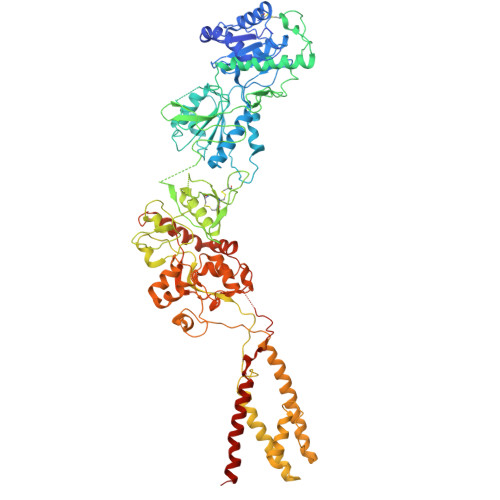
Structural basis for channel gating and blockade in tri-heteromeric GluN1-2B-2D NMDA receptor.
Kang, H., Epstein, M., Banke, T.G., Perszyk, R., Simorowski, N., Paladugu, S., Liotta, D.C., Traynelis, S.F., Furukawa, H.(2025) Neuron 113: 991
- PubMed: 39954679 Search on PubMed
- DOI: https://doi.org/10.1016/j.neuron.2025.01.013
- Primary Citation Related Structures:
9D37, 9D38, 9D39, 9D3A, 9D3B, 9D3C - PubMed Abstract:
Discrete activation of N-methyl-D-aspartate receptor (NMDAR) subtypes by glutamate and the co-agonist glycine is fundamental to neuroplasticity. A distinct variant, the tri-heteromeric receptor, comprising glycine-binding GluN1 and two types of glutamate-binding GluN2 subunits, exhibits unique pharmacological characteristics, notably enhanced sensitivity to the anti-depressant channel blocker S-(+)-ketamine. Despite its significance, the structural mechanisms underlying ligand gating and channel blockade of tri-heteromeric NMDARs remain poorly understood. Here, we identify and characterize tri-heteromeric GluN1-2B-2D NMDAR in the adult brain, resolving its structures in the activated, inhibited, and S-(+)-ketamine-blocked states. These structures reveal the ligand-dependent conformational dynamics that modulate the tension between the extracellular domain and transmembrane channels, governing channel gating and blockade. Additionally, we demonstrate that the inhibitor (S)-DQP-997-74 selectively decouples linker tension in GluN2D, offering insights into subtype-selective targeting for cognitive modulation.
- W.M. Keck Structural Biology Laboratory, Cold Spring Harbor Laboratory, Cold Spring Harbor, NY 11724, USA.
Organizational Affiliation: