The Structure of the DNA-binding Domain of the Class III ETS-Family Member SpiD from Raja eglanteria
Terrell, J.R., Vernon, T.N., Poon, G.M.K.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| SpiD | C [auth F], F [auth E] | 107 | Rostroraja eglanteria | Mutation(s): 0 | 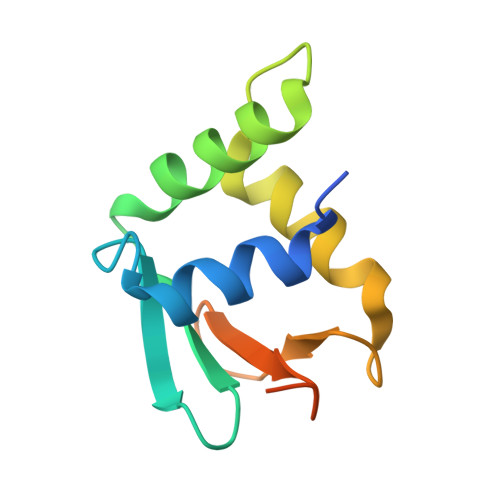 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9DEW5 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 1 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA (5'-D(*AP*AP*TP*AP*AP*AP*AP*GP*GP*AP*AP*GP*TP*GP*GP*G)-3') | A [auth C], D [auth A] | 16 | synthetic construct |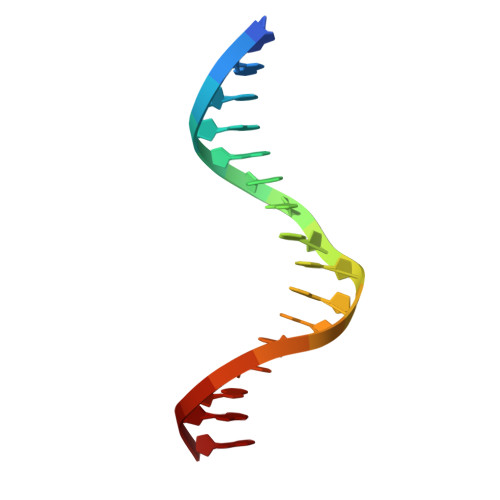  |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
Entity ID: 2 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA (5'-D(*TP*CP*CP*CP*AP*CP*TP*TP*CP*CP*TP*TP*TP*TP*AP*T)-3') | B [auth D], E [auth B] | 16 | synthetic construct |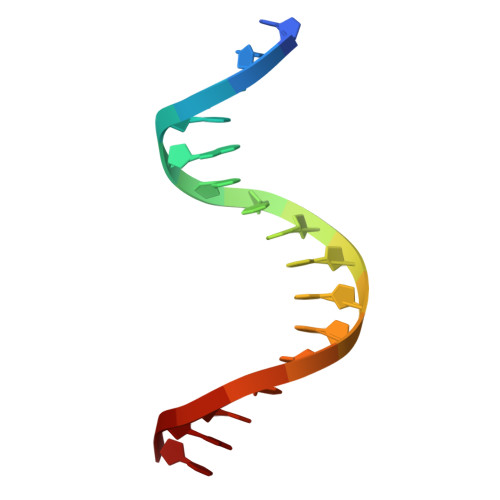  |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| EPE Download:Ideal Coordinates CCD File | G [auth B] | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID C8 H18 N2 O4 S JKMHFZQWWAIEOD-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 36.934 | α = 79.209 |
| b = 42.939 | β = 81.529 |
| c = 62.534 | γ = 76.185 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Heart, Lung, and Blood Institute (NIH/NHLBI) | United States | HL155178 |
| National Science Foundation (NSF, United States) | United States | MCB2028902 |
| National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) | United States | GM137160 |