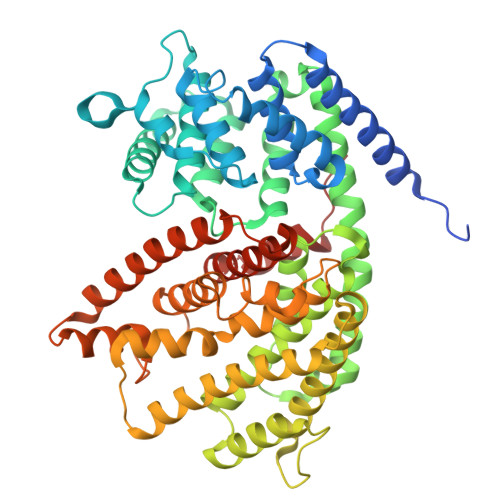
Structure and Function of Sabinene Synthase, a Monoterpene Cyclase That Generates a Highly Strained [3.1.0] Bicyclic Product.
Gaynes, M.N., Osika, K.R., Christianson, D.W.(2024) Biochemistry 63: 3147-3159
- PubMed: 39527408 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.4c00476
- Primary Citation Related Structures:
9D28, 9D29, 9D2A - PubMed Abstract:
Sabinene is a plant natural product with a distinctive strained [3.1.0] bicyclic ring system that is used commercially as a spicy and pine-like fragrance with citrus undertones. This unusual monoterpene has also been studied as an antifungal and anti-inflammatory agent as well as a next-generation biofuel. In order to understand the molecular determinants of [3.1.0] bicyclic ring formation in sabinene biosynthesis, we now report three X-ray crystal structures of sabinene synthase from Western red cedar, Thuja plicata (TpSS), with open and partially closed active site conformations at 2.21-2.72 Å resolution. We additionally report the complete biochemical characterization of sabinene synthase, including steady-state kinetics, active site mutagenesis, and product array profiling. The catalytic metal ion requirement is unexpectedly broad for a class I terpene cyclase: optimal catalytic activity was measured using Mn 2+ or Co 2+ , with more modest activity observed using Mg 2+ or Ni 2+ . Kinetic parameters were determined for both full-length TpSS and a deletion variant lacking the putative N-terminal plastidial targeting sequence, designated ΔTpSS. Monoterpene product profiles for both indicated similar product arrays independent of the catalytic metal ion used, with sabinene comprising nearly 90% of the total products generated. Site-directed mutagenesis was utilized to probe the function of active site residues, and several mutants yielded altered product arrays. Most notably, the G458A substitution converted ΔTpSS into a high-activity α-pinene synthase. α-Pinene contains a bicyclic [3.1.1] ring system; structural and mechanistic analyses suggest a molecular rationale for the reprogrammed transannulation reaction, leading to the alternative bicyclic product.
- Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania, 231 South 34th Street, Philadelphia, Pennsylvania 19104-6323, United States.
Organizational Affiliation: