Structural and immunological characterization of the H3 influenza hemagglutinin during antigenic drift.
de Paiva Froes Rocha, R., Tomris, I., Bowman, C.A., Stevens, E., Kantorow, J., Plitt, C.M., Peng, W., Oeverdieck, S., Galdino Andrade, T., Ferguson, J.A., Jung, D.D., Marques, R.E., Herfst, S., Snijder, J., Chakraborty, S., Torrents de la Pena, A., Berndsen, Z.T., de Vries, R.P., Ward, A.B.(2025) Nat Commun 16: 11452-11452
- PubMed: 41381528 Search on PubMed
- DOI: https://doi.org/10.1038/s41467-025-66375-7
- Primary Citation Related Structures:
9CXT, 9CXU, 9D0Y, 9D1U, 9D2M - PubMed Abstract:
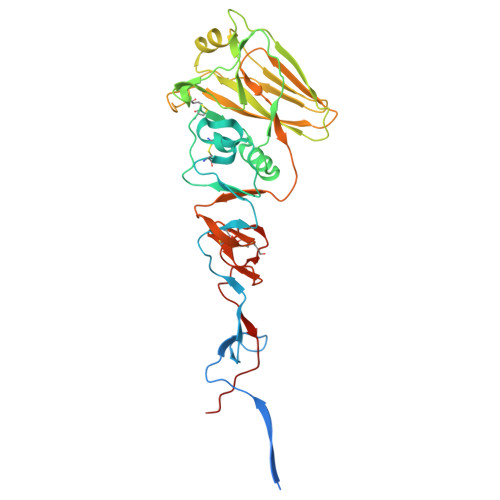
The quest for a universal influenza vaccine holds great promise for mitigating the global burden of influenza-related morbidity and mortality. However, challenges persist in identifying conserved epitopes capable of eliciting robust and durable immune responses. In this study, we explore the influence of glycan evolution on H3 hemagglutinin from 1968 to present day and its impacts on protein structure, antigenicity and immunogenicity by using computational, biochemical and biophysical techniques. Structural characterization of HK/68 and Sing/16 by cryo-electron microscopy shows that while HK/68 is resistant to enzymatic deglycosylation, removal of glycans destabilizes the hyperglycosylated head and membrane-proximal region in Sing/16. Furthermore, the appearance of glycans in Sing/16 hemagglutinin head domain shifts the polyclonal immune response upon vaccination to target the esterase and stem. These insights expand our understanding of glycans beyond their role in protein folding and highlight the interplay among glycan integration and immune recognition to design a universal influenza vaccine.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA, USA.
Organizational Affiliation: