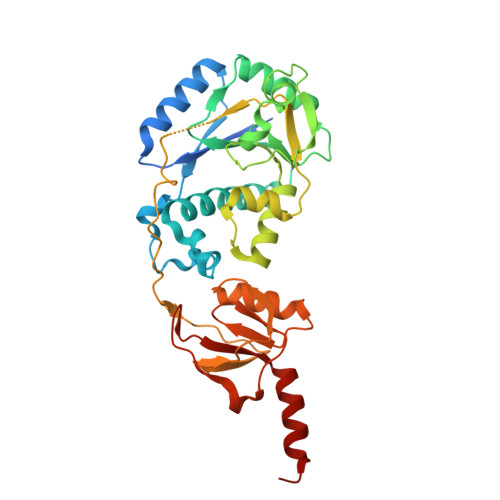
The Crystal Structure of Acanthamoeba Polyphaga Mimivirus R655 and Its Potential Glycosaminoglycan Glycosyltransferase Activity.
Kim, J.S., Zhang, B., Morin, K.H., Gilliam, M., Guo, H.(2025) Biochemistry 64: 2737-2744
- PubMed: 40545624 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.5c00058
- Primary Citation Related Structures:
9CRZ - PubMed Abstract:
Acanthamoeba polyphaga mimivirus (APMV) is a giant virus that encodes over 1,000 genes, many of which are involved in protein post-translational modifications, such as glycosylation. Here, we present the high-resolution crystal structure of a putative mimiviral glycosyltransferase R655 (Uniprot accession ID: Q5UQ62). R655 consists of a Rossmann domain with a glycosyltransferase family A (GT-A) fold followed by a CPW-WPC domain, connected by a long loop. The N-terminal region of this loop crosses over the Mn 2+ and uridine diphosphate (UDP) binding pocket in the Rossmann domain, suggesting that it may function as a pocket cap. Within the binding pocket, the aspartate residue in the EXD motif coordinates Mn 2+ while the glutamate residue is positioned to interact with the sugar moiety of the UDP-sugar. UDP is sandwiched between Leu11 and an α helix, the diphosphate of which is stabilized by a nearby Arg17. R655 contains two strips of electropositive surfaces near the Mn 2+ /UDP binding site, suggesting that it may recognize negatively charged substrates. Structural comparisons of R655 with homologous proteins in the AlphaFold Protein Structure Database and the Worldwide Protein Data Bank identified top matches, including collagen galactosyltransferases and the Vaccinia virus H3 envelope protein, a heparan sulfate-binding glycosyltransferase. However, enzymatic activity assays revealed that R655 exhibits activity toward negatively charged glycosaminoglycans (GAGs) rather than collagen, consistent with the structural findings. Kinetic parameters, cofactor, and sugar donor preferences were characterized. Mutations of the critical residues in the active site and nearby loop abolish R655's enzymatic activity. These results provide novel structural insights into R655 function and suggest a new role of APMV in glycobiology.
- Department of Molecular and Cellular Biochemistry, University of Kentucky, Lexington, Kentucky 40536, United States.
Organizational Affiliation: