Structure of a left-right-handed G-quadruplex from the promoter of the NSD1 gene
Xing, E.R., Yatsunyk, L.A.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA (25-MER) | 26 | Homo sapiens | 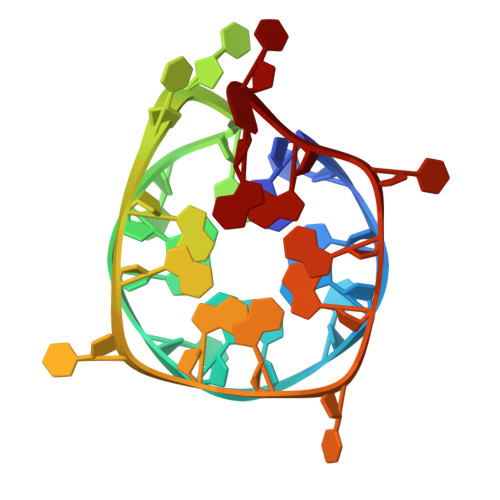 | |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| MES Download:Ideal Coordinates CCD File | I [auth A] | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID C6 H13 N O4 S SXGZJKUKBWWHRA-UHFFFAOYSA-N |  | ||
| NCO Download:Ideal Coordinates CCD File | F [auth A], G [auth A], H [auth A] | COBALT HEXAMMINE(III) Co H18 N6 DYLMFCCYOUSRTK-UHFFFAOYSA-N |  | ||
| PEG Download:Ideal Coordinates CCD File | J [auth A] K [auth A] L [auth A] M [auth A] N [auth A] | DI(HYDROXYETHYL)ETHER C4 H10 O3 MTHSVFCYNBDYFN-UHFFFAOYSA-N |  | ||
| K Download:Ideal Coordinates CCD File | B [auth A], C [auth A], D [auth A], E [auth A] | POTASSIUM ION K NPYPAHLBTDXSSS-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 71.849 | α = 90 |
| b = 71.849 | β = 90 |
| c = 41.637 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| Coot | model building |
| autoPROC | data reduction |
| autoPROC | data scaling |
| PHENIX | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Cancer Institute (NIH/NCI) | United States | 1R15CA253134 |