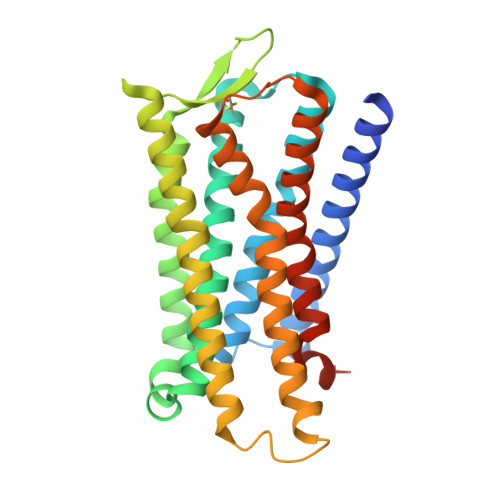
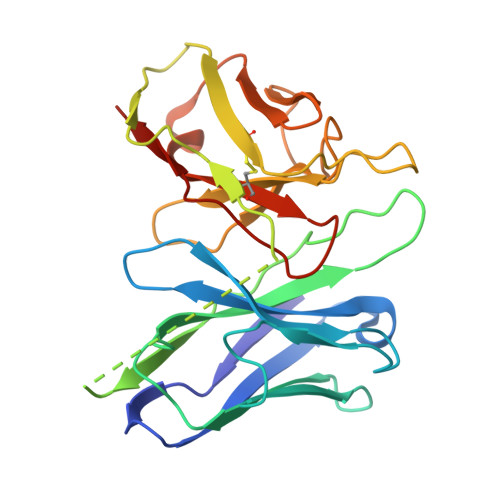
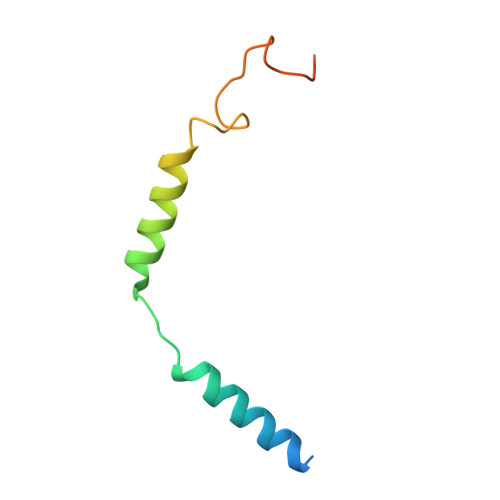
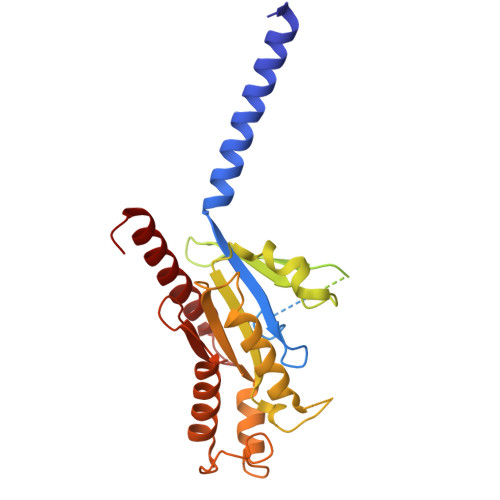
Structure-guided design of partial agonists at an opioid receptor.
Varga, B.R., Bernhard, S.M., El Daibani, A., Zaidi, S.A., Lam, J.H., Aguilar, J., Appourchaux, K., Nazarova, A.L., Kouvelis, A., Shinouchi, R., Hammond, H.R., Eans, S.O., Weinreb, V., Margolis, E.B., Fay, J.F., Huang, X.P., Pradhan, A., Katritch, V., McLaughlin, J.P., Majumdar, S., Che, T.(2025) Nat Commun 16: 2518-2518
- PubMed: 40082451 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-57734-5
- Primary Citation Related Structures:
9CGJ, 9CGK - PubMed Abstract:
Chronic pain and opioid overdose deaths highlight the need for non-addictive analgesics with novel mechanisms. The δ opioid receptor (δOR) is a promising target, as it lacks the respiratory depression associated with µ opioid receptor (µOR) agonists. However, early δOR full agonists caused seizures, limiting their clinical use. Partial δOR agonists may offer more controlled receptor activation than full agonists, but their development has been hindered by uncertainty regarding the molecular mechanism of partial agonism. Here we show that C6-Quino, a bitopic ligand developed through structure-based design, acts as a selective δOR partial agonist. Functional studies reveal that C6-Quino shows differential activity at G-protein and arrestin pathways and interacts with the sodium binding pocket, confirmed through cryo-EM analysis. C6-Quino demonstrates oral activity, analgesic activity in chronic pain models without causing δOR-related seizures and µOR-related adverse effects which have limited opioid usage in recent times. This discovery outlines a new strategy for developing δOR-targeted analgesics and provides a framework for optimizing signaling profiles of other Class A GPCRs.
- Center for Clinical Pharmacology, Department of Anesthesiology, Washington University School of Medicine, St. Louis, MO, USA.
Organizational Affiliation: