HEPN-AbiV is an RNase in the AbiV antiphage system
Zhu, X., Morency, C., Picard, M.-E., Mosterd, C., McAlister, J.A., Perrault-Jolicoeur, A., Geddes-McAlister, J., Shi, R., Moineau, S.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| AbiV family abortive infection protein | 184 | Lactococcus cremoris | Mutation(s): 0 | 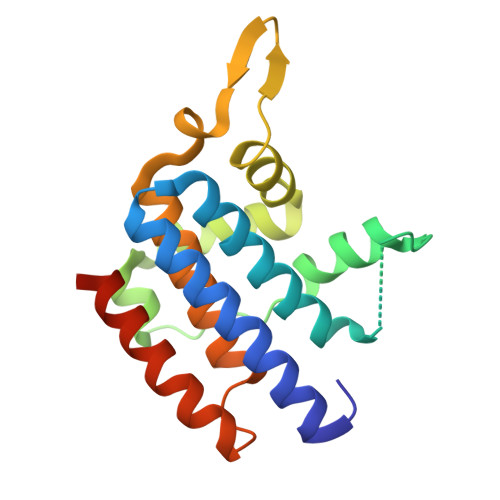 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A2RJ64 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| SO4 Download:Ideal Coordinates CCD File | AA [auth E] BA [auth G] CA [auth G] DA [auth Q] EA [auth R] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 74.444 | α = 90.96 |
| b = 106.901 | β = 89.95 |
| c = 146.228 | γ = 95.02 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| SCALA | data scaling |
| XDS | data reduction |
| MOLREP | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Natural Sciences and Engineering Research Council (NSERC, Canada) | Canada | 436202 |