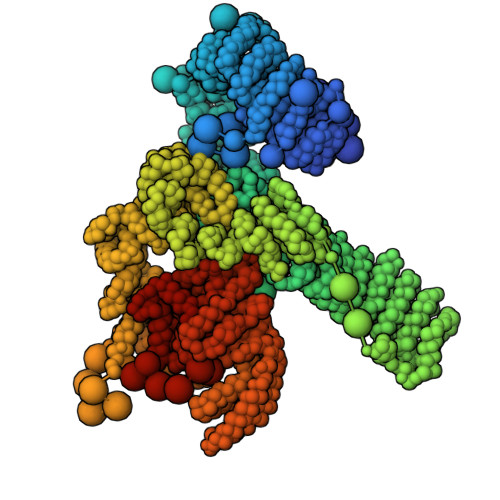
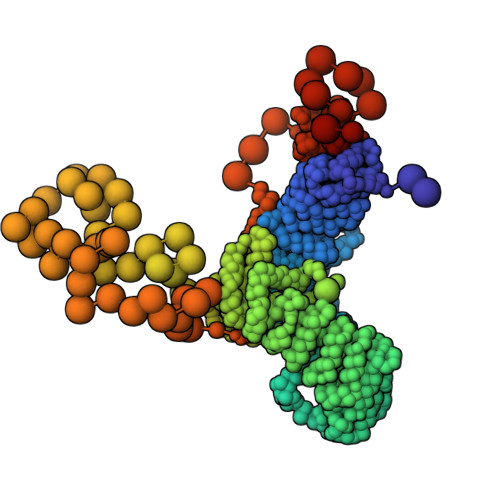
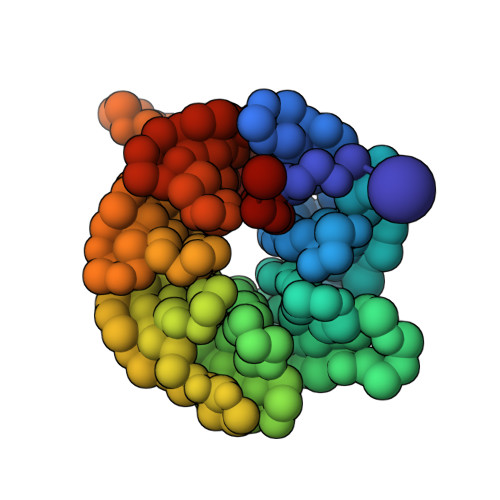
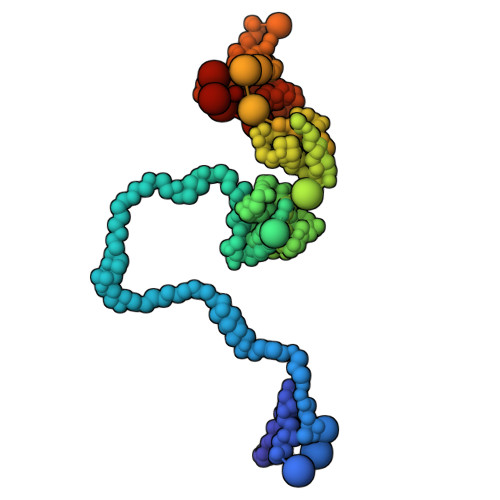
Structural Basis for the Recruitment and Selective Phosphorylation of Akt by mTORC2
Taylor, M.S., Chen, M., Hancock, M., Wranik, M., Miller, B.D., O'Meara, T.R., Palanski, B.A., Ficarro, S.B., Groendyke, B.J., Xiang, Y., Kondo, K.T., Linde-Garelli, K.Y., Lee, M.J., Mondal, D., Freund, D., Congreve, S., Matas, K., Hennik, M., Xibinaku, K., Valenstein, M.L., van Euwen, T., Marto, J.A., Sali, A., Shi, Y., Gray, N.S., Sabatini, D.M., Chu, N., Rogala, K.B., Cole, P.A.(2025) Science 391
- PubMed: 41308123
- DOI: https://doi.org/10.1126/science.adv7111
- Primary Citation Related Structures:
9AAA, 9ZBJ, 9ZBK - PubMed Abstract:
The mechanistic target of rapamycin (mTOR) protein kinase forms two multiprotein complexes, mTORC1 and mTORC2, that function in distinct signaling pathways. mTORC1 is regulated by nutrients, and mTORC2 is a central node in phosphoinositide-3 kinase (PI3K) and small guanosine triphosphate Ras signaling networks commonly deregulated in cancer and diabetes. Although mTOR phosphorylates many substrates in vitro, in cells, mTORC1 and mTORC2 have high specificity: mTORC2 phosphorylates the protein kinases Akt and PKC, but not closely related kinases that are mTORC1 substrates. To understand how mTORC2 recognizes substrates, we created semisynthetic probes to trap the mTORC2 :: Akt complex and determine its structure. Whereas most protein kinases recognize amino acids adjacent to the phosphorylation site, local sequence contributes little to substrate recognition by mTORC2. Instead, the specificity determinants were secondary and tertiary structural elements of Akt that bound the mTORC2 component mSin1 distal to the mTOR active site and were conserved among at least 18 related substrates. These results reveal how mTORC2 recognizes its canonical substrates and may enable the design of mTORC2-specific inhibitors.
- Department of Pathology and Laboratory Medicine, Brown University, Providence, RI, USA.
Organizational Affiliation: