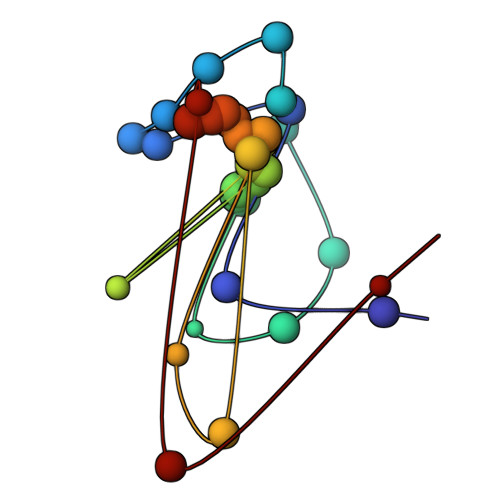
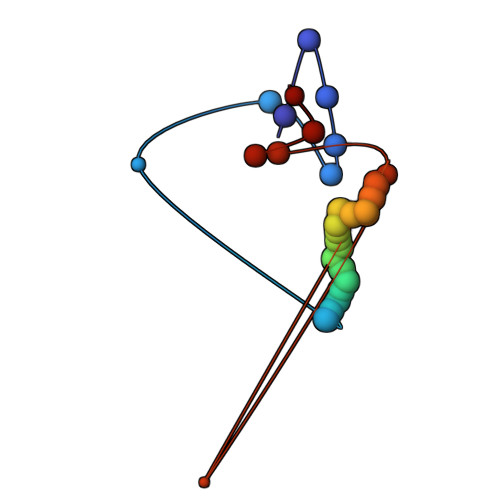
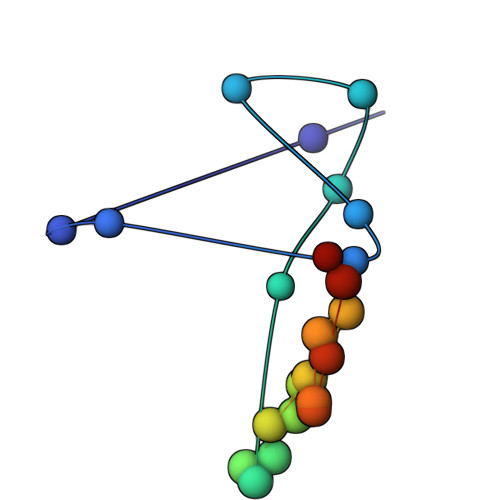
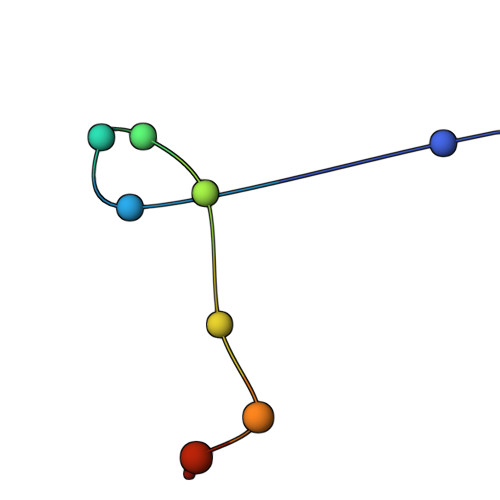
The molecular architecture of the desmosomal outer dense plaque by integrative structural modeling
Pasani, S,, Menon, K.S., Shruthi, V.(2024) Protein Sci 33
- PubMed: 39548826
- DOI: https://doi.org/10.1002/pro.5217
- Primary Citation of Related Structures:
9A8U - PubMed Abstract:
Desmosomes mediate cell-cell adhesion and are prevalent in tissues under mechanical stress. However, their detailed structural characterization is not available. Here, we characterized the molecular architecture of the desmosomal outer dense plaque (ODP) using Bayesian integrative structural modeling via the Integrative Modeling Platform. Starting principally from the structural interpretation of a cryo-electron tomography (cryo-ET) map of the ODP, we integrated information from x-ray crystallography, an immuno-electron microscopy study, biochemical assays, in silico predictions of transmembrane and disordered regions, homology modeling, and stereochemistry information. The integrative structure was validated by information from imaging, tomography, and biochemical studies that were not used in modeling. The ODP resembles a densely packed cylinder with a plakophilin (PKP) layer and a plakoglobin (PG) layer; the desmosomal cadherins and PKP span these two layers. Our integrative approach allowed us to localize disordered regions, such as the N-terminus of PKP and the C-terminus of PG. We refined previous protein-protein interactions between desmosomal proteins and provided possible structural hypotheses for defective cell-cell adhesion in several diseases by mapping disease-related mutations on the structure. Finally, we point to features of the structure that could confer resilience to mechanical stress. Our model provides a basis for generating experimentally verifiable hypotheses on the structure and function of desmosomal proteins in normal and disease states.
- National Center for Biological Sciences, Tata Institute of Fundamental Research, Bengaluru, India.
Organizational Affiliation: