Characterization of an A3G-Vif HIV-1-CRL5-CBFb structure using a cross-linking mass spectrometry pipeline for integrative modeling of host-pathogen complexes
Robyn M Kaake, Ignacia Echeverria, Seung Joong Kim, John Von Dollen, Nicholas M Chesarino, Yuqing Feng, Clinton Yu, Hai Ta, Linda Chelico, Lan Huang, John Gross, Andrej Sali, Nevan J Krogan(2021) Mol Cell Proteomics 20: 100132
- PubMed: 34389466 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.mcpro.2021.100132
- Primary Citation Related Structures:
9A1I, 9A1J - PubMed Abstract:
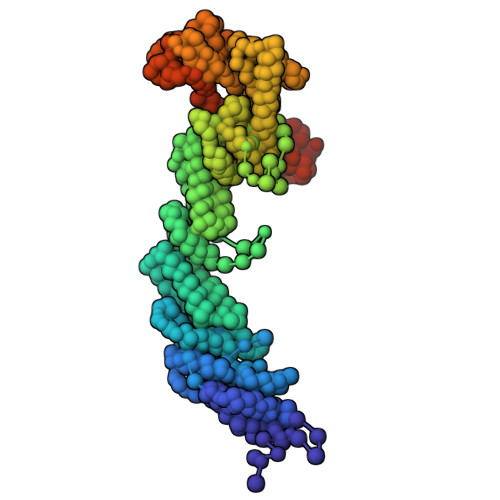
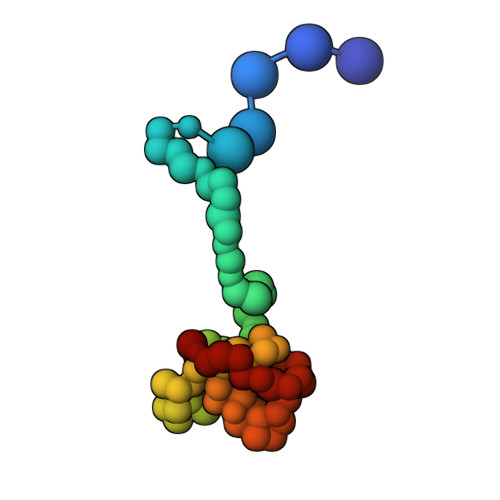
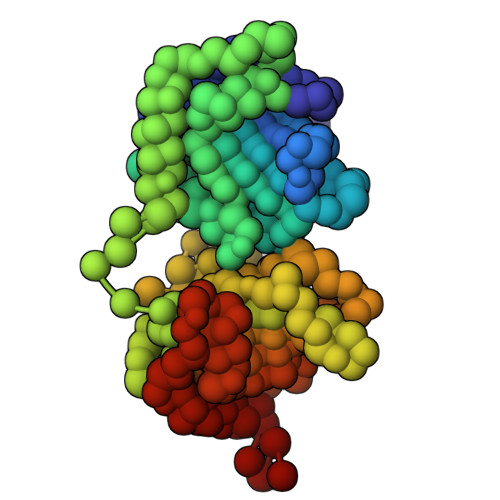
Structural analysis of host-pathogen protein complexes remains challenging, largely due to their structural heterogeneity. Here, we describe a pipeline for the structural characterization of these complexes using integrative structure modeling based on chemical cross-links and residue-protein contacts inferred from mutagenesis studies. We used this approach on the HIV-1 Vif protein bound to restriction factor APOBEC3G (A3G), the Cullin-5 E3 ring ligase (CRL5), and the cellular transcription factor Core Binding Factor Beta (CBFβ) to determine the structure of the (A3G-Vif-CRL5-CBFβ) complex. Using the MS-cleavable DSSO cross-linker to obtain a set of 132 cross-links within this reconstituted complex along with the atomic structures of the subunits and mutagenesis data, we computed an integrative structure model of the heptameric A3G-Vif-CRL5-CBFβ complex. The structure, which was validated using a series of tests, reveals that A3G is bound to Vif mostly through its N-terminal domain. Moreover, the model ensemble quantifies the dynamic heterogeneity of the A3G C-terminal domain and Cul5 positions. Finally, the model was used to rationalize previous structural, mutagenesis and functional data not used for modeling, including information related to the A3G-bound and unbound structures as well as mapping functional mutations to the A3G-Vif interface. The experimental and computational approach described here is generally applicable to other challenging host-pathogen protein complexes.
- Department of Cellular and Molecular Pharmacology, California Institute for Quantitative Biosciences, University of California, San Francisco, San Francisco, California, USA; Quantitative Biosciences Institute, University of California, San Francisco, San Francisco, California, USA; Gladstone Institute of Data Science and Biotechnology, J. David Gladstone Institutes, San Francisco, California, USA.
Organizational Affiliation: