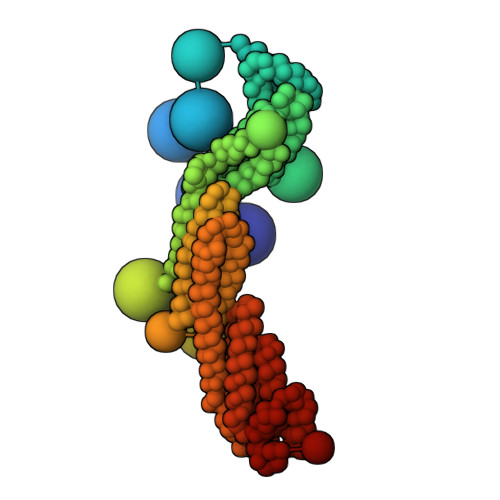
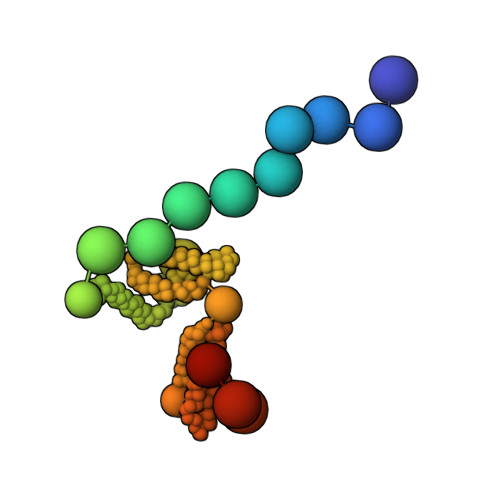
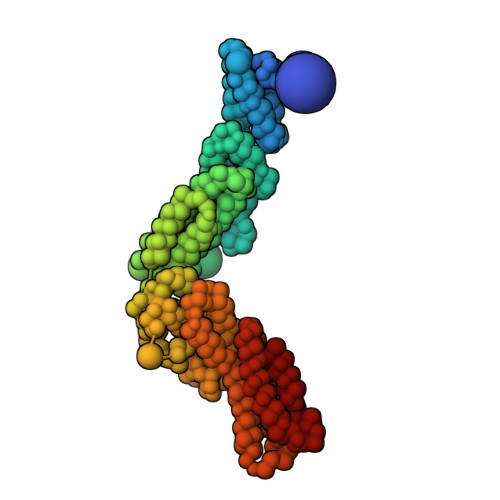
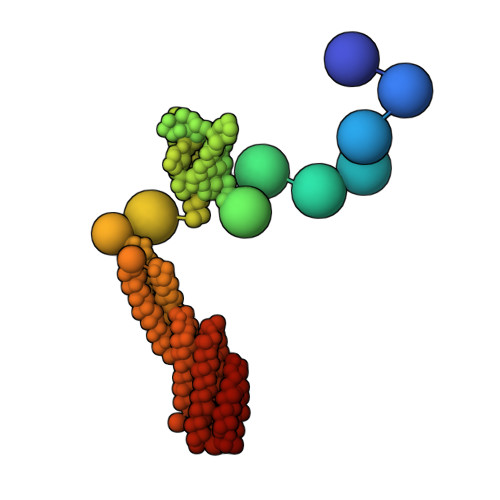
Integrative structure and function of the yeast exocyst complex.
Ganesan SJ, Feyder MJ, Chemmama IE, Fang F, Rout MP, Chait BT, Shi Y, Munson M, Sali A(2020) Protein Sci 29: 1486-1501
- PubMed: 32239688 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3863
- Primary Citation Related Structures:
9A05 - PubMed Abstract:
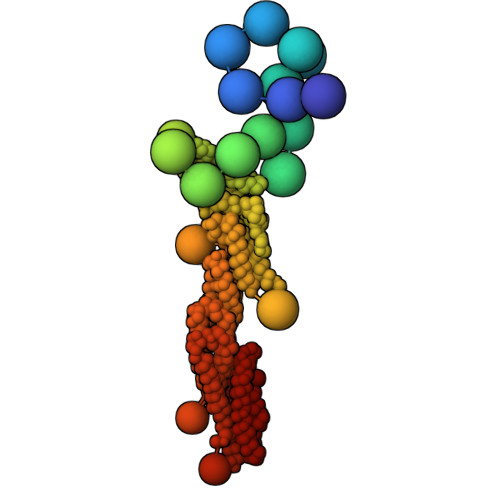
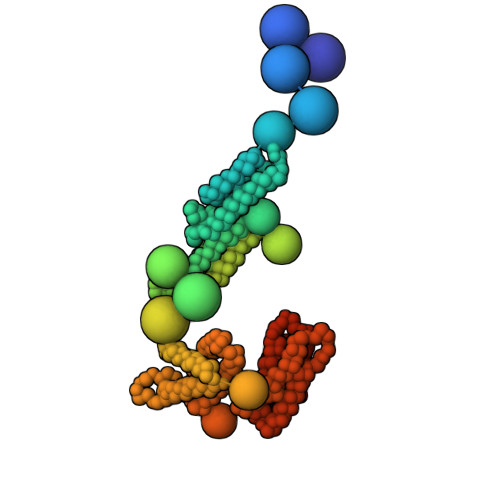
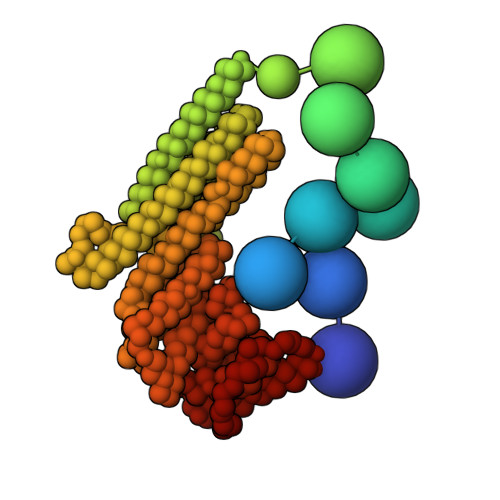
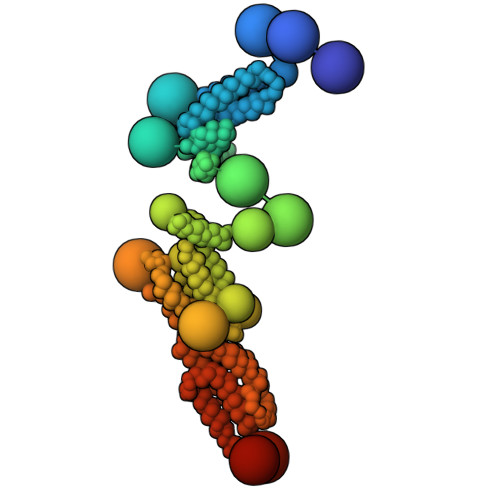
Exocyst is an evolutionarily conserved hetero-octameric tethering complex that plays a variety of roles in membrane trafficking, including exocytosis, endocytosis, autophagy, cell polarization, cytokinesis, pathogen invasion, and metastasis. Exocyst serves as a platform for interactions between the Rab, Rho, and Ral small GTPases, SNARE proteins, and Sec1/Munc18 regulators that coordinate spatial and temporal fidelity of membrane fusion. However, its mechanism is poorly described at the molecular level. Here, we determine the molecular architecture of the yeast exocyst complex by an integrative approach, based on a 3D density map from negative-stain electron microscopy (EM) at ~16 Å resolution, 434 disuccinimidyl suberate and 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide hydrochloride cross-links from chemical-crosslinking mass spectrometry, and partial atomic models of the eight subunits. The integrative structure is validated by a previously determined cryo-EM structure, cross-links, and distances from in vivo fluorescence microscopy. Our subunit configuration is consistent with the cryo-EM structure, except for Sec5. While not observed in the cryo-EM map, the integrative model localizes the N-terminal half of Sec3 near the Sec6 subunit. Limited proteolysis experiments suggest that the conformation of Exo70 is dynamic, which may have functional implications for SNARE and membrane interactions. This study illustrates how integrative modeling based on varied low-resolution structural data can inform biologically relevant hypotheses, even in the absence of high-resolution data.
- Department of Bioengineering and Therapeutic Sciences, University of California, San Francisco, San Francisco, California, USA.
Organizational Affiliation: