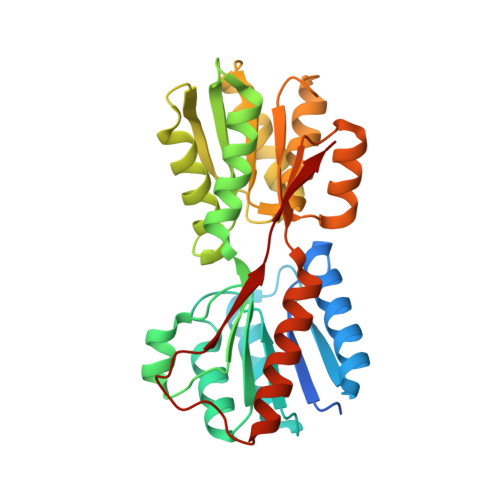
X-ray structures of Enterobacter cloacae allose-binding protein in complexes with monosaccharides demonstrate its unique recognition mechanism for high affinity to allose.
Kamitori, S.(2023) Biochem Biophys Res Commun 682: 187-192
- PubMed: 37820454
- DOI: https://doi.org/10.1016/j.bbrc.2023.10.016
- Primary Citation Related Structures:
8WL5, 8WL7, 8WL9, 8WLB - PubMed Abstract:
d-Allose is an aldohexose of the C3-epimer of d-glucose, existing in very small amounts in nature, called a rare sugar. The operon responsible for d-allose metabolism, the allose operon, was found in several bacteria, which consists of seven genes: alsR, alsB, alsA, alsC, alsE, alsK, and rpiB. To understand the biological implication of the allose operon utilizing a rare sugar of d-allose as a carbon source, it is important to clarify whether the allose operon functions specifically for d-allose or also functions for other ligands. It was proposed that the allose operon can function for d-ribose, which is essential as a component of nucleotides and abundant in nature. Allose-binding protein, AlsB, coded in the allose operon, is thought to capture a ligand outside the cell, and is expected to show high affinity for the specific ligand. X-ray structure determinations of Enterobacter cloacae AlsB (EtcAlsB) in ligand-free form, and in complexes with d-allose, d-ribose, and d-allulose, and measurements of the thermal parameters of the complex formation using an isothermal titration calorimeter were performed. The results demonstrated that EtcAlsB has a unique recognition mechanism for high affinity to d-allose by changing its conformation from an open to a closed form depending on d-allose-binding, and that the binding of d-ribose to EtcAlsB could not induce a completely closed form but an intermediate form, explaining the low affinity for d-ribose.
- Research Facility Center for Science & Technology and Faculty of Medicine, International Institute of Rare Sugar Research and Education, Kagawa University, 1750-1 Ikenobe, Miki-cho, Kita-gun, Kagawa, 761-0793, Japan. Electronic address: kamitori.shigehiro@kagawa-u.ac.jp.
Organizational Affiliation: