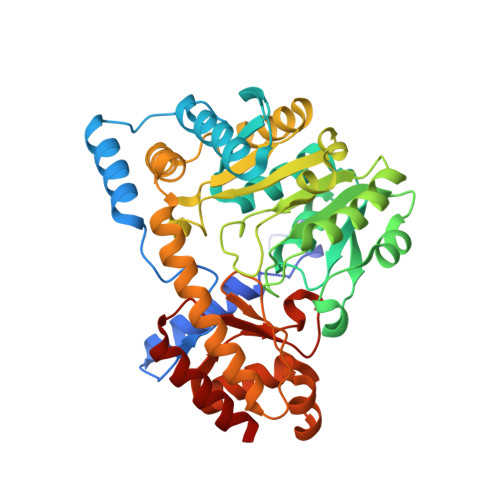
Crystal structure of an aspartate aminotransferase Lpg0070 from Legionella pneumophila.
Gao, Y., Yang, X., Hua, L., Wang, M., Ge, Q., Wang, W., Wang, N., Ma, J., Ge, H.(2023) Biochem Biophys Res Commun 689: 149230-149230
- PubMed: 37984176
- DOI: https://doi.org/10.1016/j.bbrc.2023.149230
- Primary Citation of Related Structures:
8WKJ, 8WOU - PubMed Abstract:
Legionella pneumophila aspartate aminotransferase (Lpg0070) is a member of the transaminase and belongs to the pyridoxal 5'-phosphate (PLP)-dependent superfamily. It is responsible for the transfer of α-amino between aspartate and α-ketoglutarate to form glutamate and oxaloacetate. Here, we report the crystal structure of Lpg0070 at the resolution of 2.14 Å and 1.7 Å, in apo-form and PLP-bound, respectively. Our structural analysis revealed the specific residues involved in the PLP binding and free form against PLP-bound supported conformational changes before substrate recognition. In vitro enzyme activity proves that the absence of the N-terminal arm reduces the enzyme activity of Lpg0070. These data provide further evidence to support the N-terminal arm plays a crucial role in catalytic activity.
- Institutes of Material Science and Information Technology, Anhui University, Hefei, 230601, China; School of Resources and Environmental Engineering, Anhui University, Hefei, 230601, China.
Organizational Affiliation: