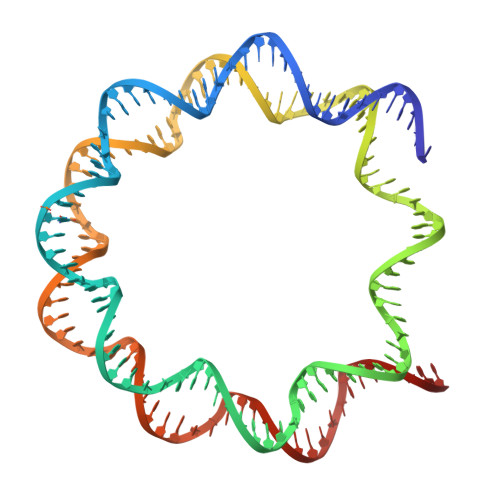
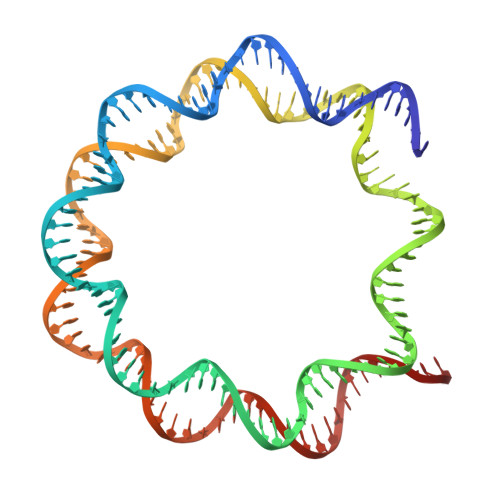
Contributing factors to the oxidation-induced mutational landscape in human cells.
Cordero, C., Mehta, K.P.M., Weaver, T.M., Ling, J.A., Freudenthal, B.D., Cortez, D., Roberts, S.A.(2024) Nat Commun 15: 10722-10722
- PubMed: 39715760 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-55497-z
- Primary Citation Related Structures:
8VWS, 8VWT, 8VWU, 8VWV - PubMed Abstract:
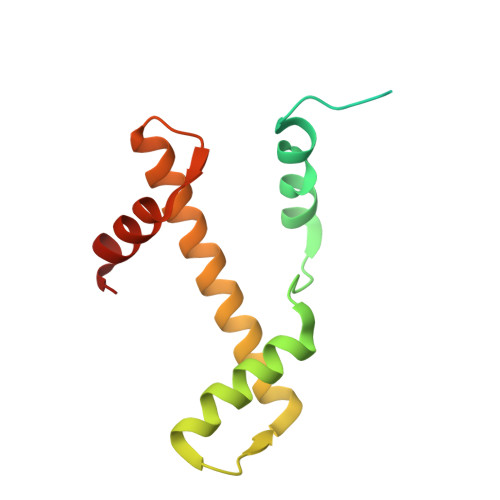
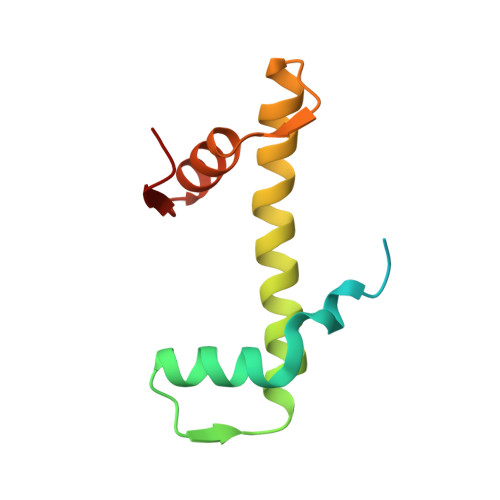
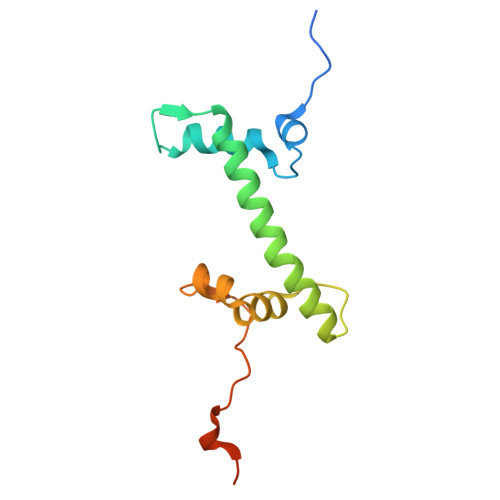
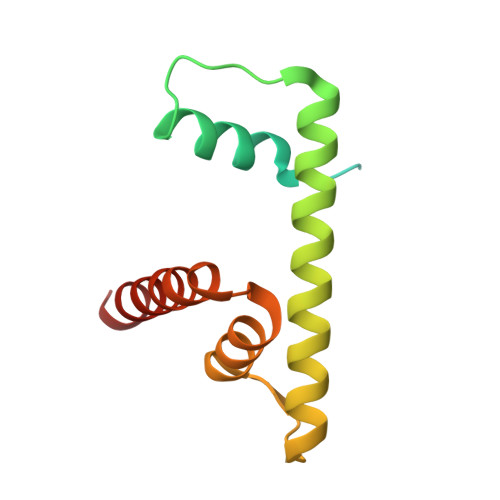
8-oxoguanine (8-oxoG) is a common oxidative DNA lesion that causes G > T substitutions. Determinants of local and regional differences in 8-oxoG-induced mutability across genomes are currently unknown. Here, we show DNA oxidation induces G > T substitutions and insertion/deletion (INDEL) mutations in human cells and cancers. Potassium bromate (KBrO 3 )-induced 8-oxoGs occur with similar sequence preferences as their derived substitutions, indicating that the reactivity of specific oxidants dictates mutation sequence specificity. While 8-oxoG occurs uniformly across chromatin, 8-oxoG-induced mutations are elevated in compact genomic regions, within nucleosomes, and at inward facing guanines within strongly positioned nucleosomes. Cryo-electron microscopy structures of OGG1-nucleosome complexes indicate that these effects originate from OGG1's ability to flip outward positioned 8-oxoG lesions into the catalytic pocket while inward facing lesions are occluded by the histone octamer. Mutation spectra from human cells with DNA repair deficiencies reveals contributions of a DNA repair network limiting 8-oxoG mutagenesis, where OGG1- and MUTYH-mediated base excision repair is supplemented by the replication-associated factors Pol η and HMCES. Transcriptional asymmetry of KBrO 3 -induced mutations in OGG1- and Pol η-deficient cells also demonstrates transcription-coupled repair can prevent 8-oxoG-induced mutation. Thus, oxidant chemistry, chromatin structures, and DNA repair processes combine to dictate the oxidative mutational landscape in human genomes.
- Department of Microbiology and Molecular Genetics, University of Vermont, Burlington, VT, 05405, USA.
Organizational Affiliation: