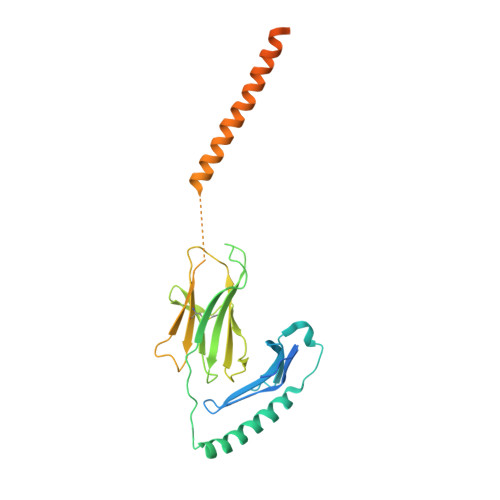
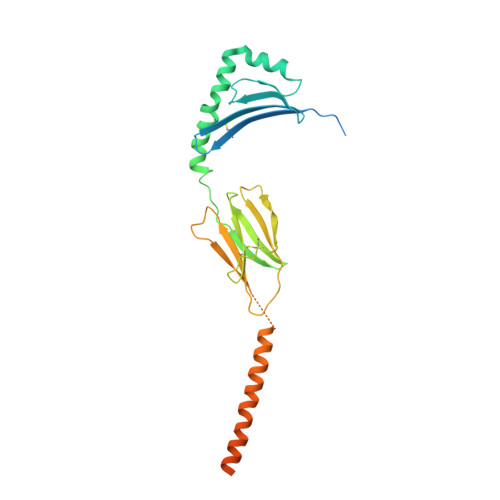
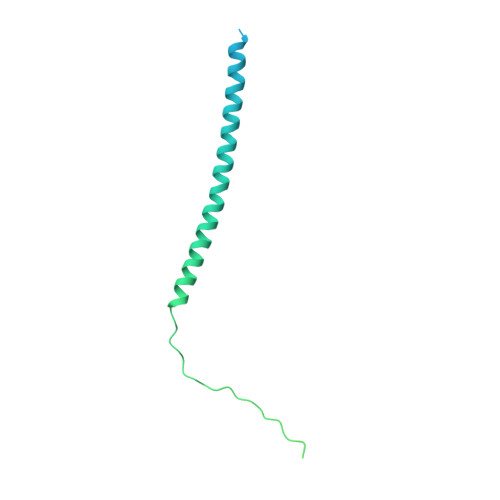
Structural insights into human MHC-II association with invariant chain.
Wang, N., Waghray, D., Caveney, N.A., Jude, K.M., Garcia, K.C.(2024) Proc Natl Acad Sci U S A 121: e2403031121-e2403031121
- PubMed: 38687785 Search on PubMed
- DOI: https://doi.org/10.1073/pnas.2403031121
- Primary Citation Related Structures:
8VRW, 8VSP - PubMed Abstract:
The loading of processed peptides on to major histocompatibility complex II (MHC-II) molecules for recognition by T cells is vital to cell-mediated adaptive immunity. As part of this process, MHC-II associates with the invariant chain (Ii) during biosynthesis in the endoplasmic reticulum to prevent premature peptide loading and to serve as a scaffold for subsequent proteolytic processing into MHC-II-CLIP. Cryo-electron microscopy structures of full-length Human Leukocyte Antigen-DR (HLA-DR) and HLA-DQ complexes associated with Ii, resolved at 3.0 to 3.1 Å, elucidate the trimeric assembly of the HLA/Ii complex and define atomic-level interactions between HLA, Ii transmembrane domains, loop domains, and class II-associated invariant chain peptides (CLIP). Together with previous structures of MHC-II peptide loading intermediates DO and DM, our findings complete the structural path governing class II antigen presentation.
- Department of Molecular and Cellular Physiology, Stanford University School of Medicine, Stanford, CA 94305.
Organizational Affiliation: