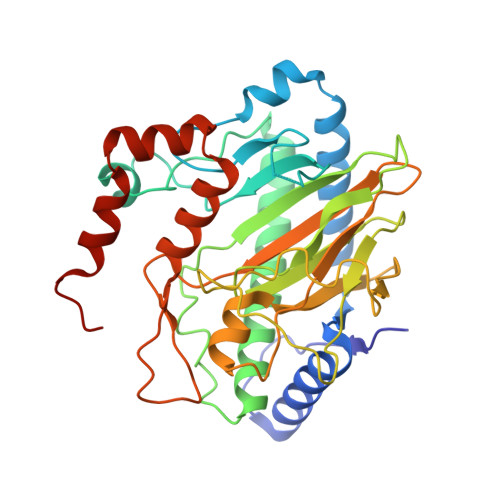
Structural, Spectroscopic, and Computational Insights from Canavanine-Bound and Two Catalytically Compromised Variants of the Ethylene-Forming Enzyme.
Chatterjee, S., Fellner, M., Rankin, J., Thomas, M.G., J S Rifayee, S.B., Christov, C.Z., Hu, J., Hausinger, R.P.(2024) Biochemistry 63: 1038-1050
- PubMed: 38577885 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.4c00031
- Primary Citation Related Structures:
6CBA, 6CF3, 8UC2 - PubMed Abstract:
The ethylene-forming enzyme (EFE) is an Fe(II), 2-oxoglutarate (2OG), and l-arginine (l-Arg)-dependent oxygenase that either forms ethylene and three CO 2 /bicarbonate from 2OG or couples the decarboxylation of 2OG to C5 hydroxylation of l-Arg. l-Arg binds with C5 toward the metal center, causing 2OG to change from monodentate to chelate metal interaction and OD1 to OD2 switch of D191 metal coordination. We applied anaerobic UV-visible spectroscopy, X-ray crystallography, and computational approaches to three EFE systems with high-resolution structures. The ineffective l-Arg analogue l-canavanine binds to the EFE with O5 pointing away from the metal center while promoting chelate formation by 2OG but fails to switch the D191 metal coordination from OD1 to OD2. Substituting alanine for R171 that interacts with 2OG and l-Arg inactivates the protein, prevents metal chelation by 2OG, and weakens l-Arg binding. The R171A EFE had electron density at the 2OG binding site that was identified by mass spectrometry as benzoic acid. The substitution by alanine of Y306 in the EFE, a residue 12 Å away from the catalytic metal center, generates an interior cavity that leads to multiple local and distal structural changes that reduce l-Arg binding and significantly reduce the enzyme activity. Flexibility analyses revealed correlated and anticorrelated motions in each system, with important distinctions from the wild-type enzyme. In combination, the results are congruent with the currently proposed enzyme mechanism, reinforce the importance of metal coordination by OD2 of D191, and highlight the importance of the second coordination sphere and longer range interactions in promoting EFE activity.
- Department of Microbiology, Genetics, and Immunology, Michigan State University, East Lansing, Michigan 48824, United States.
Organizational Affiliation: