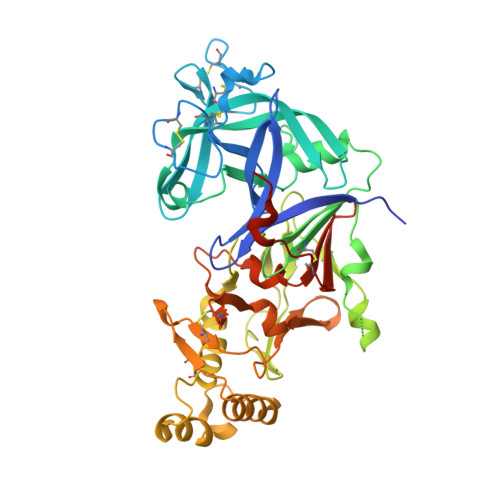
Structure-activity analysis of imino-pyrimidinone-fused pyrrolidines aids the development of dual plasmepsin V and plasmepsin X inhibitors.
Hodder, A.N., Sleebs, B.E., Adams, G., Rezazadeh, S., Ngo, A., Jarman, K., Scally, S., Czabotar, P., Wang, H., McCauley, J.A., Olsen, D.B., Cowman, A.F.(2025) FEBS J 292: 2843-2864
- PubMed: 40035447 Search on PubMed
- DOI: https://doi.org/10.1111/febs.70038
- Primary Citation Related Structures:
8TYF, 8TYG, 8TYH - PubMed Abstract:
A library of known aspartic protease inhibitors was screened to identify compounds that inhibit plasmepsin V from Plasmodium vivax. This screen revealed compounds with an imino-pyrimidinone-fused pyrrolidine (IPF) scaffold that exhibited sub-micromolar inhibitory activity against plasmepsin V. Further screening of IPF analogs against the related aspartic protease plasmepsin X showed inhibitory activity, while a third aspartic protease, plasmepsin IX, was not significantly inhibited. Modifications to the P1 biaryl region of the IPF scaffold differentially modulated inhibition of both plasmepsin V and X. Notably, analogs with potent plasmepsin X inhibitory activity successfully blocked the growth of Plasmodium falciparum in vitro. X-ray structures of IPF analogs in complex with plasmepsin V provided insights into their binding mode and revealed avenues to further improve IPF potency and selectivity between plasmepsin V and X. This understanding of how these compounds interact with the active sites of plasmepsin V and X will serve as a foundation for the future design of dual inhibitors targeting these proteases.
- The Walter and Eliza Hall Institute of Medical Research, Parkville, Australia.
Organizational Affiliation: