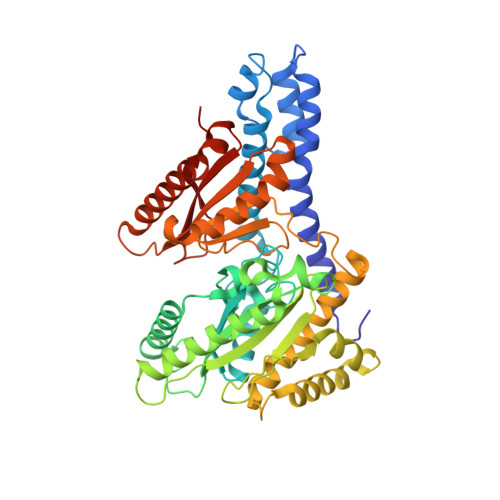
Cyanobacterial alpha-carboxysome carbonic anhydrase is allosterically regulated by the Rubisco substrate RuBP.
Pulsford, S.B., Outram, M.A., Forster, B., Rhodes, T., Williams, S.J., Badger, M.R., Price, G.D., Jackson, C.J., Long, B.M.(2024) Sci Adv 10: eadk7283-eadk7283
- PubMed: 38728392 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.adk7283
- Primary Citation Related Structures:
8THM - PubMed Abstract:
Cyanobacterial CO 2 concentrating mechanisms (CCMs) sequester a globally consequential proportion of carbon into the biosphere. Proteinaceous microcompartments, called carboxysomes, play a critical role in CCM function, housing two enzymes to enhance CO 2 fixation: carbonic anhydrase (CA) and Rubisco. Despite its importance, our current understanding of the carboxysomal CAs found in α-cyanobacteria, CsoSCA, remains limited, particularly regarding the regulation of its activity. Here, we present a structural and biochemical study of CsoSCA from the cyanobacterium Cyanobium sp. PCC7001. Our results show that the Cyanobium CsoSCA is allosterically activated by the Rubisco substrate ribulose-1,5-bisphosphate and forms a hexameric trimer of dimers. Comprehensive phylogenetic and mutational analyses are consistent with this regulation appearing exclusively in cyanobacterial α-carboxysome CAs. These findings clarify the biologically relevant oligomeric state of α-carboxysomal CAs and advance our understanding of the regulation of photosynthesis in this globally dominant lineage.
- ARC Centre of Excellence in Synthetic Biology, Sydney, NSW, Australia.
Organizational Affiliation: