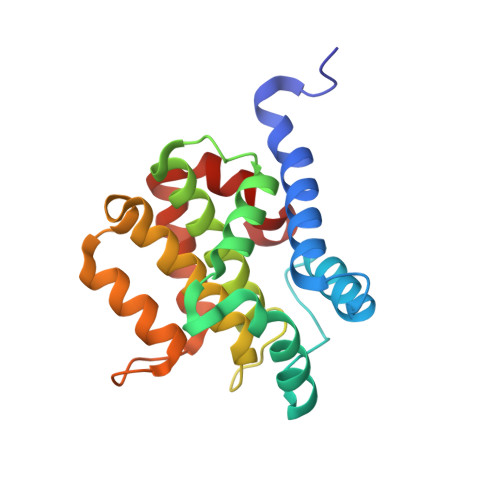
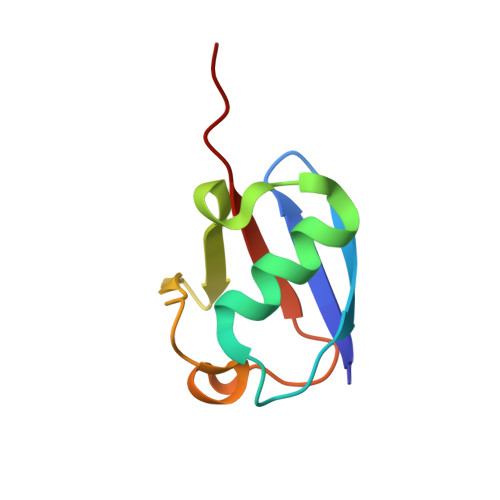
Bacterial ligases reveal fundamental principles of polyubiquitin specificity.
Franklin, T.G., Brzovic, P.S., Pruneda, J.N.(2023) Mol Cell 83: 4538-4554.e4
- PubMed: 38091999 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2023.11.017
- Primary Citation Related Structures:
8ST7, 8ST8, 8ST9 - PubMed Abstract:
Homologous to E6AP C terminus (HECT) E3 ubiquitin (Ub) ligases direct substrates toward distinct cellular fates dictated by the specific form of monomeric or polymeric Ub (polyUb) signal attached. How polyUb specificity is achieved has been a long-standing mystery, despite extensive study in various hosts, ranging from yeast to human. The bacterial pathogens enterohemorrhagic Escherichia coli and Salmonella Typhimurium encode outlying examples of "HECT-like" (bHECT) E3 ligases, but commonalities to eukaryotic HECT (eHECT) mechanism and specificity had not been explored. We expanded the bHECT family with examples in human and plant pathogens. Three bHECT structures in primed, Ub-loaded states resolved key details of the entire Ub ligation process. One structure provided a rare glimpse into the act of ligating polyUb, yielding a means to rewire polyUb specificity of both bHECT and eHECT ligases. Studying this evolutionarily distinct bHECT family has revealed insight into the function of key bacterial virulence factors as well as fundamental principles underlying HECT-type Ub ligation.
- Department of Molecular Microbiology and Immunology, Oregon Health and Science University, Portland, OR 97239, USA.
Organizational Affiliation: