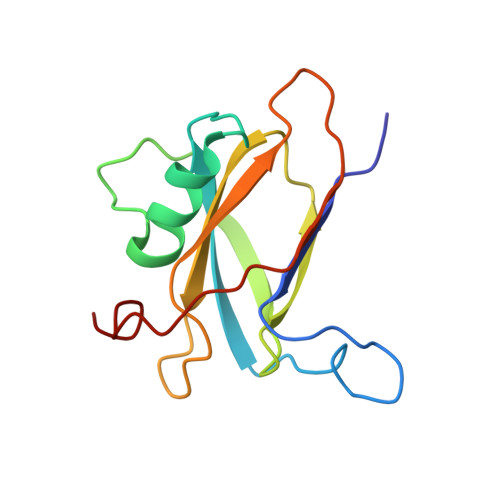
The basal and major pilins in the Corynebacterium diphtheriae SpaA pilus adopt similar structures that competitively react with the pilin polymerase.
Sue, C.K., Cheung, N.A., Mahoney, B.J., McConnell, S.A., Scully, J.M., Fu, J.Y., Chang, C., Ton-That, H., Loo, J.A., Clubb, R.T.(2024) Biopolymers 115: e23539-e23539
- PubMed: 37227047 Search on PubMed
- DOI: https://doi.org/10.1002/bip.23539
- Primary Citation Related Structures:
8SQX - PubMed Abstract:
Many species of pathogenic gram-positive bacteria display covalently crosslinked protein polymers (called pili or fimbriae) that mediate microbial adhesion to host tissues. These structures are assembled by pilus-specific sortase enzymes that join the pilin components together via lysine-isopeptide bonds. The archetypal SpaA pilus from Corynebacterium diphtheriae is built by the Cd SrtA pilus-specific sortase, which crosslinks lysine residues within the SpaA and SpaB pilins to build the shaft and base of the pilus, respectively. Here, we show that Cd SrtA crosslinks SpaB to SpaA via a K139(SpaB)-T494(SpaA) lysine-isopeptide bond. Despite sharing only limited sequence homology, an NMR structure of SpaB reveals striking similarities with the N-terminal domain of SpaA ( N SpaA) that is also crosslinked by Cd SrtA. In particular, both pilins contain similarly positioned reactive lysine residues and adjacent disordered AB loops that are predicted to be involved in the recently proposed "latch" mechanism of isopeptide bond formation. Competition experiments using an inactive SpaB variant and additional NMR studies suggest that SpaB terminates SpaA polymerization by outcompeting N SpaA for access to a shared thioester enzyme-substrate reaction intermediate.
- Department of Chemistry and Biochemistry, University of California, Los Angeles, California, USA.
Organizational Affiliation: