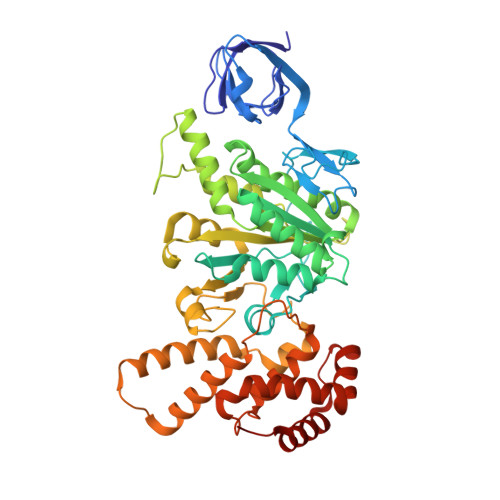
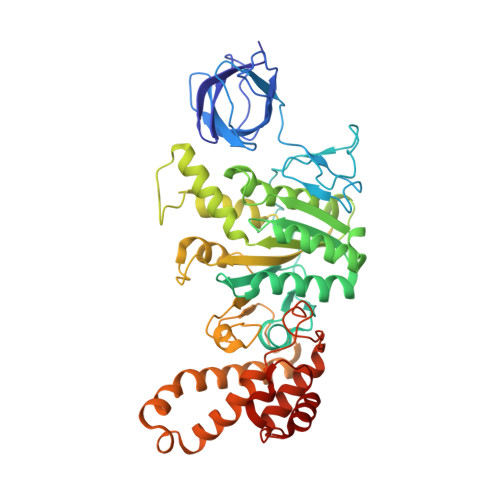
The series of conformational states adopted by rotorless F 1 -ATPase during its hydrolysis cycle.
Sobti, M., Ueno, H., Brown, S.H.J., Noji, H., Stewart, A.G.(2024) Structure 32: 393
- PubMed: 38237595 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2023.12.014
- Primary Citation Related Structures:
8SPV, 8SPW, 8SPX - PubMed Abstract:
F 1 F o ATP synthase interchanges phosphate transfer energy and proton motive force via a rotary catalytic mechanism and isolated F 1 -ATPase subcomplexes can also hydrolyze ATP to generate rotation of their central γ rotor subunit. As ATP is hydrolyzed, the F 1 -ATPase cycles through a series of conformational states that mediates unidirectional rotation of the rotor. However, even in the absence of a rotor, the α and β subunits are still able to pass through a series of conformations, akin to those that generate rotation. Here, we use cryoelectron microscopy to establish the structures of these rotorless states. These structures indicate that cooperativity in this system is likely mediated by contacts between the β subunit lever domains, irrespective of the presence of the γ rotor subunit. These findings provide insight into how long-range information may be transferred in large biological systems.
- Molecular, Structural and Computational Biology Division, The Victor Chang Cardiac Research Institute, Darlinghurst, NSW, Australia; St Vincent's Clinical School, Faculty of Medicine, UNSW Sydney, Kensington, NSW, Australia.
Organizational Affiliation: