Integrating artificial intelligence-based epitope prediction in a SARS-CoV-2 antibody discovery pipeline: caution is warranted.
Acar, D.D., Witkowski, W., Wejda, M., Wei, R., Desmet, T., Schepens, B., De Cae, S., Sedeyn, K., Eeckhaut, H., Fijalkowska, D., Roose, K., Vanmarcke, S., Poupon, A., Jochmans, D., Zhang, X., Abdelnabi, R., Foo, C.S., Weynand, B., Reiter, D., Callewaert, N., Remaut, H., Neyts, J., Saelens, X., Gerlo, S., Vandekerckhove, L.(2024) EBioMedicine 100: 104960-104960
- PubMed: 38232633 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.ebiom.2023.104960
- Primary Citation Related Structures:
8QPR, 8QQ0 - PubMed Abstract:
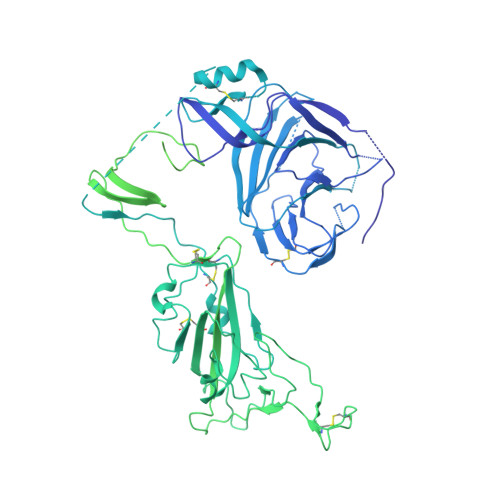
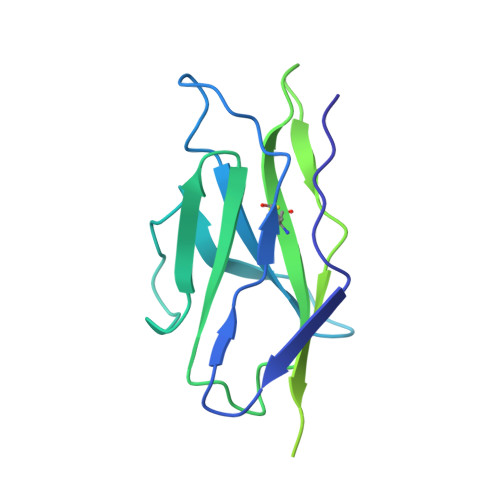
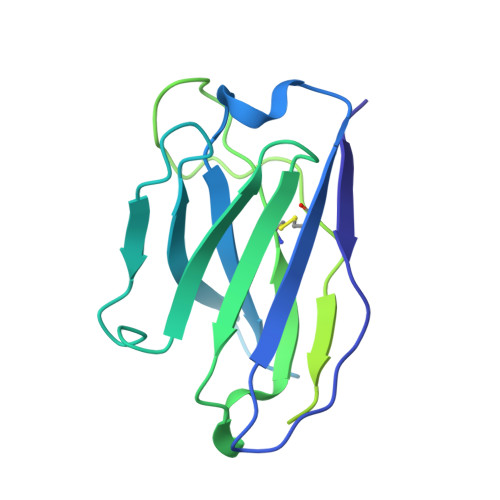
SARS-CoV-2-neutralizing antibodies (nABs) showed great promise in the early phases of the COVID-19 pandemic. The emergence of resistant strains, however, quickly rendered the majority of clinically approved nABs ineffective. This underscored the imperative to develop nAB cocktails targeting non-overlapping epitopes. Undertaking a nAB discovery program, we employed a classical workflow, while integrating artificial intelligence (AI)-based prediction to select non-competing nABs very early in the pipeline. We identified and in vivo validated (in female Syrian hamsters) two highly potent nABs. Despite the promising results, in depth cryo-EM structural analysis demonstrated that the AI-based prediction employed with the intention to ensure non-overlapping epitopes was inaccurate. The two nABs in fact bound to the same receptor-binding epitope in a remarkably similar manner. Our findings indicate that, even in the Alphafold era, AI-based predictions of paratope-epitope interactions are rough and experimental validation of epitopes remains an essential cornerstone of a successful nAB lead selection. Full list of funders is provided at the end of the manuscript.
- HIV Cure Research Center, Department of Internal Medicine and Pediatrics, Ghent University Hospital, Ghent University, Ghent 9000, Belgium.
Organizational Affiliation: