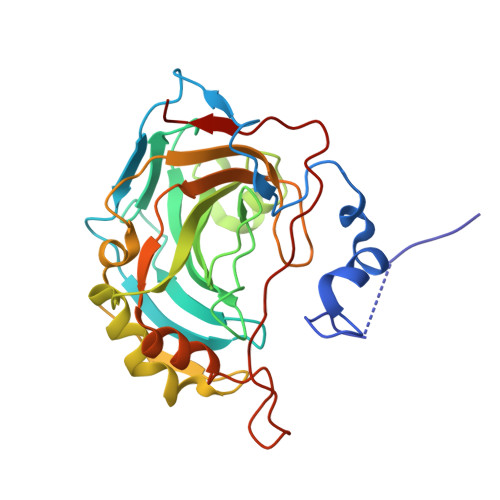
Lasamide, a Potent Human Carbonic Anhydrase Inhibitor from the Market: Inhibition Profiling and Crystallographic Studies.
Baroni, C., D'Agostino, I., Renzi, G., Kilbile, J.T., Tamboli, Y., Ferraroni, M., Carradori, S., Capasso, C., Carta, F., Supuran, C.T.(2024) ACS Med Chem Lett 15: 1749-1755
- PubMed: 39411526 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.4c00341
- Primary Citation Related Structures:
8QH8, 8QHG, 8QHJ - PubMed Abstract:
Lasamide is a synthetic precursor and a contaminant of the diuretic Furosemide manufacturing process and represents a highly valuable building block for fragment-based drug discovery approaches. We assessed the ability of Lasamide to inhibit in vitro the human-expressed Carbonic Anhydrases by means of the stopped-flow technique, and we assessed its binding modes within hCAs II and XII-mimic catalytic clefts by X-ray crystallography. Interestingly, an unprecedented crystal form for the hCA IX mimic H-tag is reported and discussed herein.
- Department of Chemistry "Ugo Schiff", University of Florence, 50019, Sesto Fiorentino, Florence, Italy.
Organizational Affiliation: