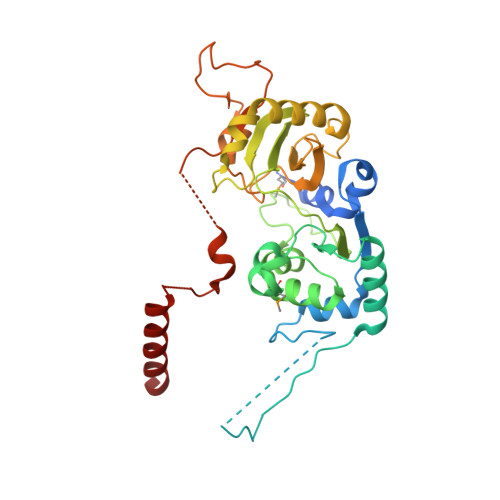
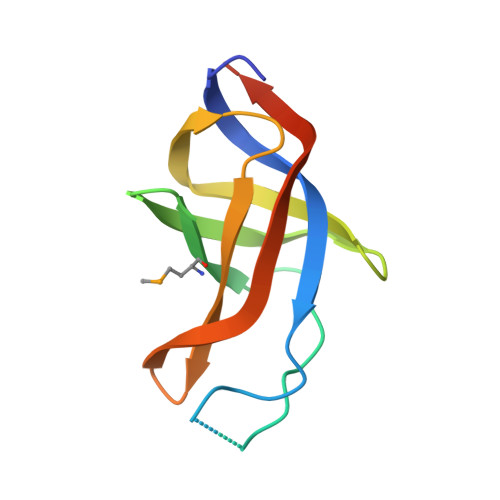
Binding of human Cdc123 to eIF2 gamma.
Cardenal Peralta, C., Vandroux, P., Neumann-Arnold, L., Panvert, M., Fagart, J., Seufert, W., Mechulam, Y., Schmitt, E.(2023) J Struct Biol 215: 108006-108006
- PubMed: 37507029 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2023.108006
- Primary Citation Related Structures:
8PHD, 8PHV - PubMed Abstract:
Eukaryotic initiation factor 2 (eIF2) plays a key role in protein synthesis and in its regulation. The assembly of this heterotrimeric factor is facilitated by Cdc123, a member of the ATP grasp family that binds the γ subunit of eIF2. Notably, some mutations related to MEHMO syndrome, an X-linked intellectual disability, affect Cdc123-mediated eIF2 assembly. The mechanism of action of Cdc123 is unclear and structural information for the human protein is awaited. Here, the crystallographic structure of human Cdc123 (Hs-Cdc123) bound to domain 3 of human eIF2γ (Hs-eIF2γD3) was determined. The structure shows that the domain 3 of eIF2γ is bound to domain 1 of Cdc123. In addition, the long C-terminal region of Hs-Cdc123 provides a link between the ATP and Hs-eIF2γD3 binding sites. A thermal shift assay shows that ATP is tightly bound to Cdc123 whereas the affinity of ADP is much smaller. Yeast cell viability experiments, western blot analysis and two-hybrid assays show that ATP is important for the function of Hs-Cdc123 in eIF2 assembly. These data and recent findings allow us to propose a refined model to explain the mechanism of action of Cdc123 in eIF2 assembly.
- Laboratoire de Biologie Structurale de la Cellule, BIOC, Ecole polytechnique, CNRS, Institut Polytechnique de Paris, 91128 Palaiseau cedex, France.
Organizational Affiliation: