C-2 Thiophenyl Tryptophan Trimers Inhibit Cellular Entry of SARS-CoV-2 through Interaction with the Viral Spike (S) Protein.
Gargantilla, M., Frances, C., Adhav, A., Forcada-Nadal, A., Martinez-Gualda, B., Marti-Mari, O., Lopez-Redondo, M.L., Melero, R., Marco-Marin, C., Gougeard, N., Espinosa, C., Rubio-Del-Campo, A., Ruiz-Partida, R., Hernandez-Sierra, M.D.P., Villamayor-Belinchon, L., Bravo, J., Llacer, J.L., Marina, A., Rubio, V., San-Felix, A., Geller, R., Perez-Perez, M.J.(2023) J Med Chem 66: 10432-10457
- PubMed: 37471688 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.3c00576
- Primary Citation Related Structures:
8P99, 8P9Y - PubMed Abstract:
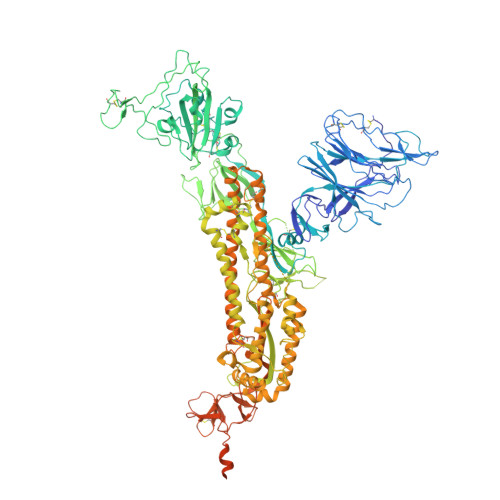
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) causes COVID-19, by infecting cells via the interaction of its spike protein (S) with the primary cell receptor angiotensin-converting enzyme (ACE2). To search for inhibitors of this key step in viral infection, we screened an in-house library of multivalent tryptophan derivatives. Using VSV-S pseudoparticles, we identified compound 2 as a potent entry inhibitor lacking cellular toxicity. Chemical optimization of 2 rendered compounds 63 and 65 , which also potently inhibited genuine SARS-CoV-2 cell entry. Thermofluor and microscale thermophoresis studies revealed their binding to S and to its isolated receptor binding domain (RBD), interfering with the interaction with ACE2. High-resolution cryoelectron microscopy structure of S, free or bound to 2 , shed light on cell entry inhibition mechanisms by these compounds. Overall, this work identifies and characterizes a new class of SARS-CoV-2 entry inhibitors with clear potential for preventing and/or fighting COVID-19.
- Instituto de Química Médica (IQM, CSIC), c/Juan de la Cierva 3, Madrid 28006, Spain.
Organizational Affiliation: