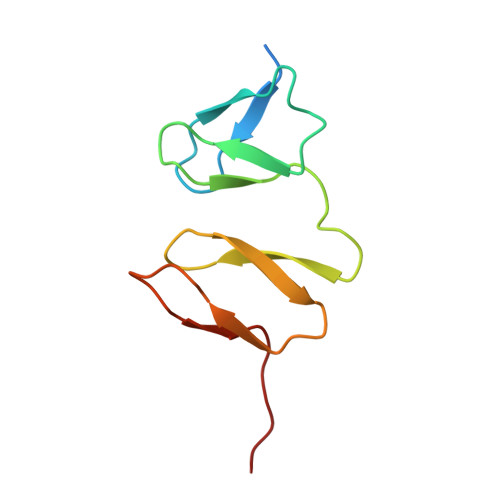
Structural basis for binding of RILPL1 to TMEM55B reveals a lysosomal platform for adaptor assembly through a conserved peptide motif.
Waschbusch, D., Pal, P., Nirujogi, R.S., Cavin, M., Singh, J., Alessi, D.R., Khan, A.R.(2026) Structure 34: 296-310.e5
- PubMed: 41314214
- DOI: https://doi.org/10.1016/j.str.2025.11.003
- Primary Citation Related Structures:
8OQH, 9QM9 - PubMed Abstract:
Inherited mutations in VPS35 and LRRK2 kinase lead to hyperphosphorylation of Rab GTPases. RH2 domain-containing proteins from the RILP homology family, such as RILPL1, are Rab effectors that recognize the LRRK2-phosphorylated switch 2 threonine of phospho-Rab8A and phospho-Rab10. Phospho-Rabs are also seen on lysosomal membranes in complex with RILPL1 and TMEM55B, a 284-residue lysosomal membrane protein lacking homology to known proteins. Here, we report crystal structures of the cytosolic region 80-166 of TMEM55B alone and in complex with a C-terminal RILPL1 peptide, which we define as the TMEM55B-binding motif (TBM). The RILPL1 TBM sits in a shallow groove across two tandem RING-like domains of TMEM55B, each forming a Zn 2+ -stabilized 40-residue β-sandwich. Co-immunoprecipitation and mass spectrometry studies indicate that TMEM55B forms complexes independently of phospho-Rabs with conserved TBMs found in JIP3, JIP4, OCRL, WDR81, and TBC1D9B. These studies suggest that TMEM55B acts as a central hub for adaptor recruitment on lysosomes.
- School of Biochemistry and Immunology, Trinity College Dublin, Dublin 2, Ireland.
Organizational Affiliation: