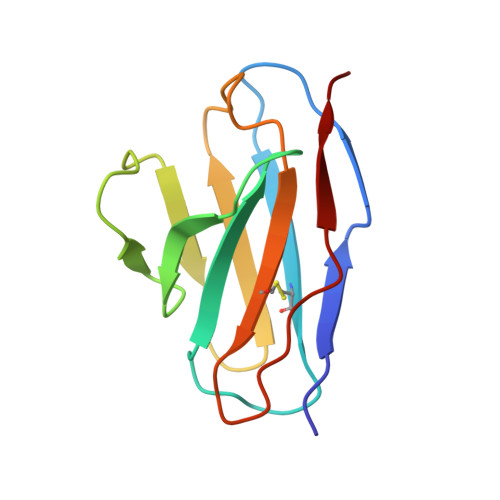
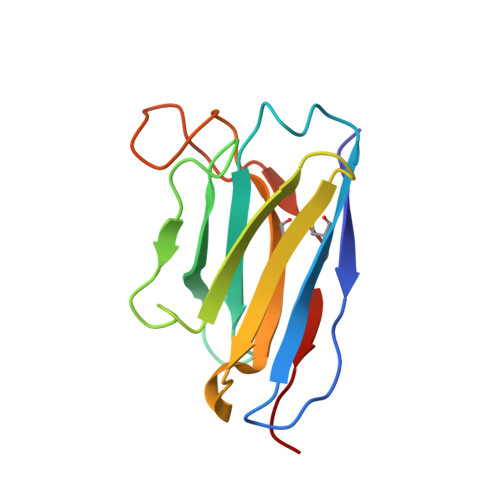
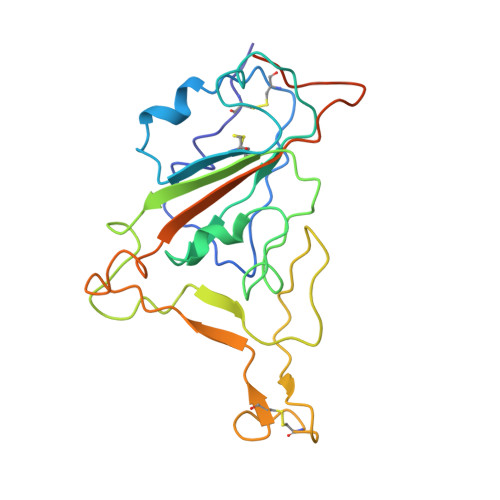
Structural basis for broad neutralization of human antibody against Omicron sublineages and evasion by XBB variant.
Sun, H., Wang, Y., Chen, X., Jiang, Y., Wang, S., Huang, Y., Liu, L., Li, Y., Lan, M., Guo, H., Yuan, Q., Zhang, Y., Li, T., Yu, H., Gu, Y., Zhang, J., Li, S., Zheng, Z., Zheng, Q., Xia, N.(2023) J Virol 97: e0113723-e0113723
- PubMed: 37855619 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/jvi.01137-23
- Primary Citation Related Structures:
8IX3 - PubMed Abstract:
The ongoing COVID-19 pandemic has been characterized by the emergence of new SARS-CoV-2 variants including the highly transmissible Omicron XBB sublineages, which have shown significant resistance to neutralizing antibodies (nAbs). This resistance has led to decreased vaccine effectiveness and therefore result in breakthrough infections and reinfections, which continuously threaten public health. To date, almost all available therapeutic nAbs, including those authorized under Emergency Use Authorization nAbs that were previously clinically useful against early strains, have recently been found to be ineffective against newly emerging variants. In this study, we provide a comprehensive structural basis about how the Class 3 nAbs, including 1G11 in this study and noted LY-CoV1404, are evaded by the newly emerged SARS-CoV-2 variants.
- State Key Laboratory of Molecular Vaccinology and Molecular Diagnostics, National Institute of Diagnostics and Vaccine Development in Infectious Diseases, School of Public Health, School of Life Sciences, Xiamen University , Xiamen, China.
Organizational Affiliation: