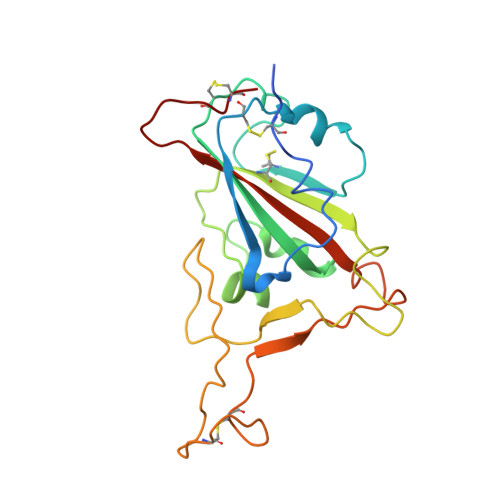
Two antibodies show broad, synergistic neutralization against SARS-CoV-2 variants by inducing conformational change within the RBD.
Sun, H., Deng, T., Zhang, Y., Lin, Y., Jiang, Y., Jiang, Y., Huang, Y., Song, S., Cui, L., Li, T., Xiong, H., Lan, M., Liu, L., Li, Y., Fang, Q., Yu, K., Jiang, W., Zhou, L., Que, Y., Zhang, T., Yuan, Q., Cheng, T., Zhang, Z., Yu, H., Zhang, J., Luo, W., Li, S., Zheng, Q., Gu, Y., Xia, N.(2024) Protein Cell 15: 121-134
- PubMed: 37470320 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/procel/pwad040
- Primary Citation Related Structures:
8IV4, 8IV5, 8IV8, 8IVA - PubMed Abstract:
Continual evolution of the severe acute respiratory syndrome coronavirus (SARS-CoV-2) virus has allowed for its gradual evasion of neutralizing antibodies (nAbs) produced in response to natural infection or vaccination. The rapid nature of these changes has incited a need for the development of superior broad nAbs (bnAbs) and/or the rational design of an antibody cocktail that can protect against the mutated virus strain. Here, we report two angiotensin-converting enzyme 2 competing nAbs-8H12 and 3E2-with synergistic neutralization but evaded by some Omicron subvariants. Cryo-electron microscopy reveals the two nAbs synergistic neutralizing virus through a rigorous pairing permitted by rearrangement of the 472-489 loop in the receptor-binding domain to avoid steric clashing. Bispecific antibodies based on these two nAbs tremendously extend the neutralizing breadth and restore neutralization against recent variants including currently dominant XBB.1.5. Together, these findings expand our understanding of the potential strategies for the neutralization of SARS-CoV-2 variants toward the design of broad-acting antibody therapeutics and vaccines.
- State Key Laboratory of Molecular Vaccinology and Molecular Diagnostics, School of Public Health, School of Life Sciences, Xiamen University, Xiamen 361102, China.
Organizational Affiliation: