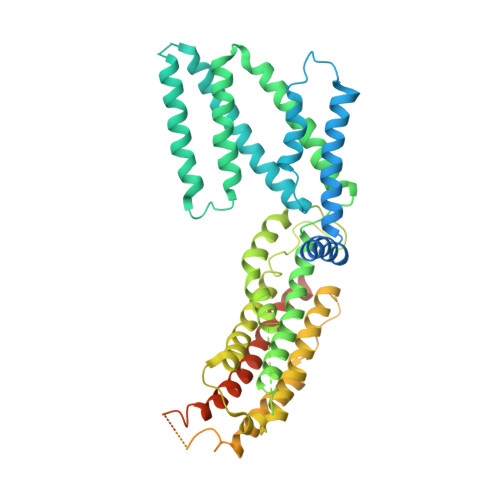
Structural insight into the Arabidopsis vacuolar anion channel ALMT9 shows clade specificity.
Qian, D., Chai, Y., Li, W., Cui, B., Lin, S., Wang, Z., Wang, C., Qu, L.Q., Gong, D.(2024) Cell Rep 43: 114731-114731
- PubMed: 39269901 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2024.114731
- Primary Citation Related Structures:
8HIW, 8HIY, 8ZVF - PubMed Abstract:
The Arabidopsis thaliana aluminum-activated malate transporter 9 (AtALMT9) functions as a vacuolar chloride channel that regulates the stomatal aperture. Here, we present the cryoelectron microscopy (cryo-EM) structures of AtALMT9 in three distinct states. AtALMT9 forms a dimer, and the pore is lined with four positively charged rings. The apo-AtALMT9 state shows a putative endogenous citrate obstructing the pore, where two W120 constriction residues enclose a gate with a pore radius of approximately 1.8 Å, representing an open state. Interestingly, channel closure is solely controlled by W120. Compared to wild-type plants, the W120A mutant exhibits more sensitivity to drought stress and is unable to restore the visual phenotype on leaves upon water recovery, reflecting persistent stomatal opening. Furthermore, notable variations are noted in channel gating and substrate recognition of Glycine max ALMT12, AtALMT9, and AtALMT1. In summary, our investigation enhances comprehension of the interplay between structure and function within the ALMT family.
- State Key Laboratory of Medicinal Chemical Biology and College of Life Sciences, Nankai University, Tianjin 300350, China. Electronic address: qiandd@nankai.edu.cn.
Organizational Affiliation: