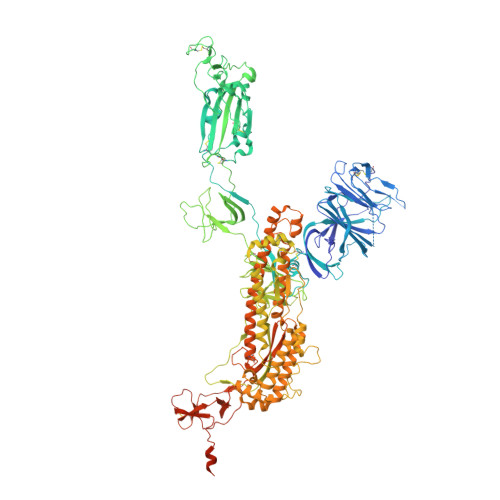
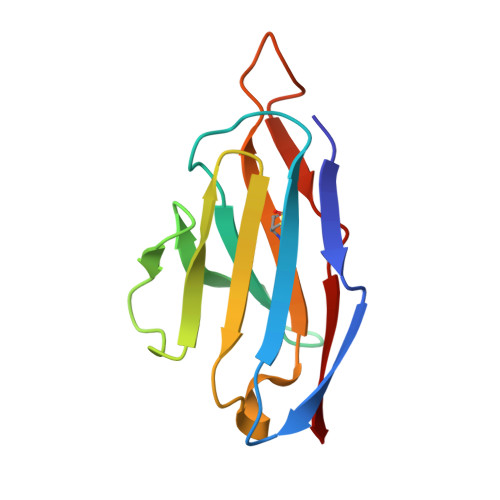
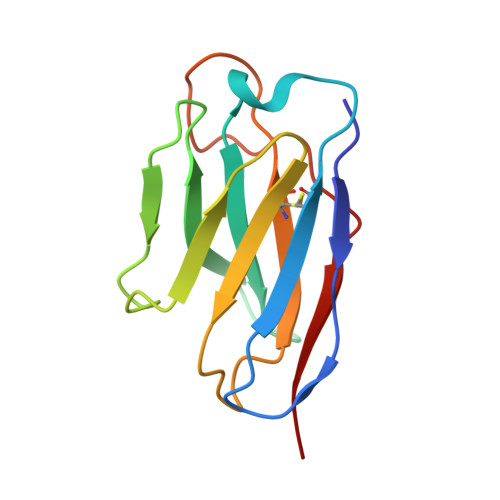
Mechanism of an RBM-targeted rabbit monoclonal antibody 9H1 neutralizing SARS-CoV-2.
Chu, X., Ding, X., Yang, Y., Lu, Y., Li, T., Gao, Y., Zheng, L., Xiao, H., Yang, T., Cheng, H., Huang, H., Liu, Y., Lou, Y., Wu, C., Chen, Y., Yang, H., Ji, X., Guo, H.(2023) Biochem Biophys Res Commun 660: 43-49
- PubMed: 37062240 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bbrc.2023.04.002
- Primary Citation Related Structures:
8HEB, 8HEC, 8HED - PubMed Abstract:
The COVID-19 pandemic, caused by SARS-CoV-2, has led to over 750 million infections and 6.8 million deaths worldwide since late 2019. Due to the continuous evolution of SARS-CoV-2, many significant variants have emerged, creating ongoing challenges to the prevention and treatment of the pandemic. Therefore, the study of antibody responses against SARS-CoV-2 is essential for the development of vaccines and therapeutics. Here we perform single particle cryo-electron microscopy (cryo-EM) structure determination of a rabbit monoclonal antibody (RmAb) 9H1 in complex with the SARS-CoV-2 wild-type (WT) spike trimer. Our structural analysis shows that 9H1 interacts with the receptor-binding motif (RBM) region of the receptor-binding domain (RBD) on the spike protein and by directly competing with angiotensin-converting enzyme 2 (ACE2), it blocks the binding of the virus to the receptor and achieves neutralization. Our findings suggest that utilizing rabbit-derived mAbs provides valuable insights into the molecular interactions between neutralizing antibodies and spike proteins and may also facilitate the development of therapeutic antibodies and expand the antibody library.
- The State Key Laboratory of Pharmaceutical Biotechnology, School of Life Sciences, Nanjing University, Nanjing, Jiangsu, 210023, China.
Organizational Affiliation: