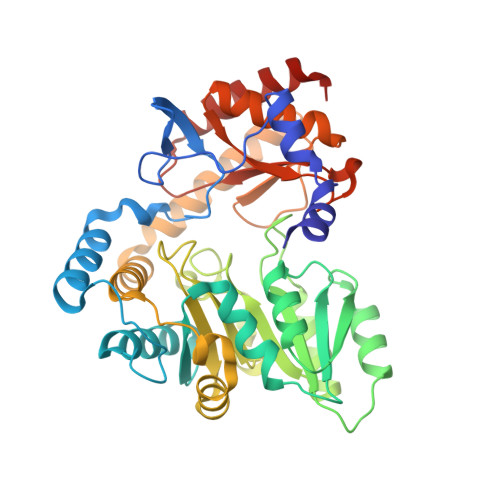
Crystal structure of Sphingobacterium multivorum serine palmitoyltransferase complexed with tris(hydroxymethyl)aminomethane.
Ikushiro, H., Takahashi, A., Murakami, T., Katayama, A., Sawai, T., Goto, H., Miyahara, I., Kamiya, N., Yano, T.(2022) Acta Crystallogr F Struct Biol Commun 78: 408-415
- PubMed: 36458620 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X22010937
- Primary Citation Related Structures:
8GUH - PubMed Abstract:
Serine palmitoyltransferase (SPT) catalyses the first reaction in sphingolipid biosynthesis: the decarboxylative condensation of L-serine (L-Ser) and palmitoyl-CoA to form 3-ketodihydrosphingosine. SPT from Sphingobacterium multivorum has been isolated and its crystal structure in complex with L-Ser has been determined at 2.3 Å resolution (PDB entry 3a2b). However, the quality of the crystal was not good enough to judge the conformation of the cofactor molecule and the orientations of the side chains of the amino-acid residues in the enzyme active site. The crystal quality was improved by revision of the purification procedure and by optimization of both the crystallization procedure and the post-crystallization treatment conditions. Here, the crystal structure of SPT complexed with tris(hydroxymethyl)aminomethane (Tris), a buffer component, was determined at 1.65 Å resolution. The protein crystallized at 20°C and diffraction data were collected from the crystals to a resolution of 1.65 Å. The crystal belonged to the tetragonal space group P4 1 2 1 2, with unit-cell parameters a = b = 61.32, c = 208.57 Å. Analysis of the crystal structure revealed C4-C5-C5A-O4P (77°) and C5-C5A-O4P-P (-143°) torsion angles in the phosphate-group moiety of the cofactor pyridoxal 5'-phosphate (PLP) that are more reasonable than those observed in the previously reported crystal structure (14° and 151°, respectively). Furthermore, the clear electron density showing a Schiff-base linkage between PLP and the bulky artificial ligand Tris indicated exceptional flexibility of the active-site cavity of this enzyme. These findings open up the possibility for further study of the detailed mechanisms of substrate recognition and catalysis by this enzyme.
- Department of Biochemistry, Faculty of Medicine, Osaka Medical and Pharmaceutical University, 2-7 Daigakumachi, Takatsuki, Osaka 569-8686, Japan.
Organizational Affiliation: