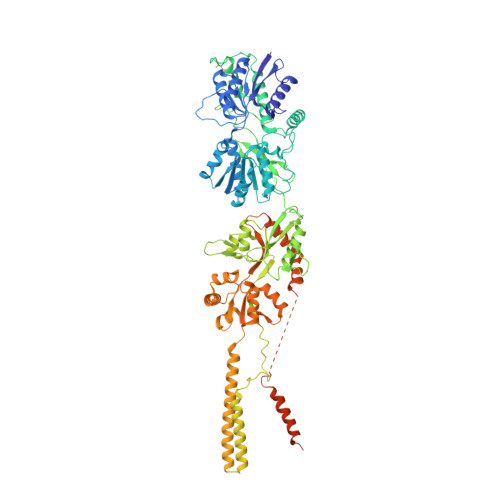
Structural basis of GluK2 kainate receptor activation by a partial agonist.
Segura-Covarrubias, G., Zhou, C., Bogdanovic, N., Zhang, L., Tajima, N.(2025) Nat Struct Mol Biol 32: 1456-1469
- PubMed: 40442317 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-025-01566-w
- Primary Citation Related Structures:
8GC5, 9C5Y, 9C5Z, 9C60, 9CAZ - PubMed Abstract:
Kainate receptors (KARs) belong to the family of ionotropic glutamate receptors that regulate neurotransmitter release and excitatory synaptic transmission in the central nervous system. Despite their critical roles in synaptic signaling and disease, the detailed gating mechanisms of KARs are not completely understood. Here we present cryo-electron microscopy structures of homomeric rat GluK2 KAR in an unliganded apo state and in complexes with a partial agonist, domoate. Partial agonist-bound GluK2 populates multiple conformations, including intermediate and desensitized states. Moreover, we demonstrate that the N-glycans at the amino-terminal domain-ligand binding domain (LBD) interface modulate receptor gating properties by interfering with cation binding at the LBD dimer interface. Together, these results provide insights into the unique gating mechanisms of KARs.
- Department of Physiology and Biophysics, Case Western Reserve University School of Medicine, Cleveland, OH, USA.
Organizational Affiliation: