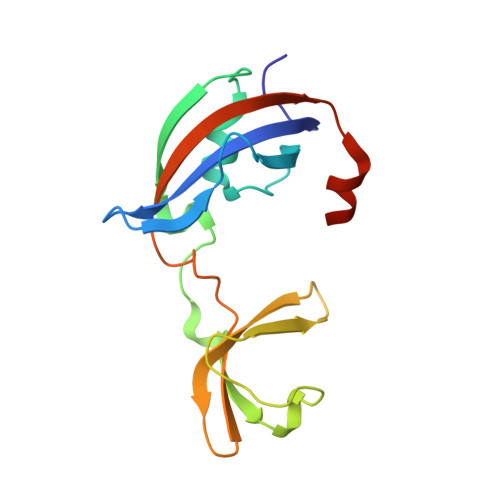
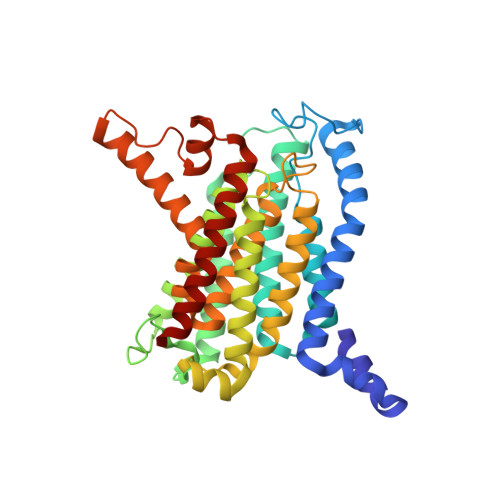
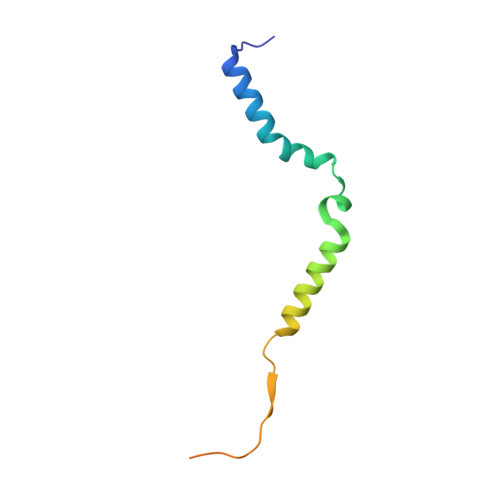
The mechanism of the phage-encoded protein antibiotic from Phi X174.
Orta, A.K., Riera, N., Li, Y.E., Tanaka, S., Yun, H.G., Klaic, L., Clemons Jr., W.M.(2023) Science 381: eadg9091-eadg9091
- PubMed: 37440661 Search on PubMed
- DOI: https://doi.org/10.1126/science.adg9091
- Primary Citation Related Structures:
8G01, 8G02 - PubMed Abstract:
The historically important phage ΦX174 kills its host bacteria by encoding a 91-residue protein antibiotic called protein E. Using single-particle electron cryo-microscopy, we demonstrate that protein E bridges two bacterial proteins to form the transmembrane YES complex [MraY, protein E, sensitivity to lysis D (SlyD)]. Protein E inhibits peptidoglycan biosynthesis by obstructing the MraY active site leading to loss of lipid I production. We experimentally validate this result for two different viral species, providing a clear model for bacterial lysis and unifying previous experimental data. Additionally, we characterize the Escherichia coli MraY structure-revealing features of this essential enzyme-and the structure of the chaperone SlyD bound to a protein. Our structures provide insights into the mechanism of phage-mediated lysis and for structure-based design of phage therapeutics.
- Division of Chemistry and Chemical Engineering, California Institute of Technology, Pasadena, CA 91125, USA.
Organizational Affiliation: